
Case Report
J Bacteriol Mycol. 2016; 3(3): 1032.
Dermatophytic Disease: Report of A Tunisian Case Caused by a Sterile Dermatophyte Identified as Trichophyton Rubrum by ITS1 and ITS4 Sequencing
Dhib I¹*, Khammari I¹, Yaacoub A¹, Gaied Meksi S¹, Letscher V², Filisetti D², Ben Said M¹ and Fathallah A¹
¹Parasitology-Mycology Laboratory, Farhat Hached Hospital, Sousse, Tunisia
²Institut of Parasitology and Tropical Pathology of Faculty of Medicine of Strasbourg, France
*Corresponding author: Imen Dhib, Faculty of Medicine, Parasitology-Mycology laboratory, Tunisia
Received: July 08, 2016; Accepted: August 22, 2016; Published: August 25, 2016
Abstract
Dermatophytic disease is a rare chronic mycosis characterized by polymorphic cutaneous lesions and possible visceral invasion, and is associated with immunodeficiency and refractoriness to drug treatment. We report herein a dermatophytic disease tunisian case caused by a sterile dermatophyte further identified as Trichophyton rubrum by ITS1 and ITS4 sequencing.
The case we report herein highlights the usefulness of molecular techniques in identification of causative agent when the mycological criteria are lacking.
Keywords: Dermatophytic disease; Sterile dermatophyte; Trichophyton rubrum; Sequencing
Introduction
Subcutaneous mycoses, which are much less common than superficial fungal infections, include a heterogeneous group of infections that often result from direct penetration of the fungus into the dermis and subcutaneous tissue through traumatic injury [1]. Dermatophytic disease is a rare chronic mycosis characterized by polymorphic cutaneous lesions and possible visceral invasion, and is associated with immunodeficiency and refractoriness to drug treatment. Dermatophytic disease has mainly described in North Africa [2]; in Tunisia, most previously reported cases are caused by Trichophyton violaceum or Microsporum canis [3,4]. The problematic identification of filamentous fungi when classical morphological criteria are missing is now feasible by DNA sequencing [5].
We report herein a dermatophytic disease Tunisian case caused by a sterile dermatophyte further identified as Trichophyton rubrum by ITS1 and ITS4 sequencing.
Case Presentation
The patient is a 43 year-old tunisian man and no known immunodeficiency. He was addressed to the lab of Mycology- Parasitology of Farhat Hached hospital, Sousse, Tunisia, for cutaneous lesions very evocative of tinea corporis. The lesions first appeared on the abdomen (Figure 1) and progressively spread to the back and the members. Later, papulo-nodules and vegetating plaques appeared and involved large parts of the body (Figure 2). On examination, neither onychomycosis nor tinea capitis were associated to skin lesions and no lymph nodes nor visceral involvement could be demonstrated.
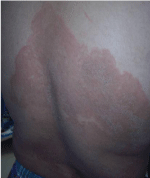
Figure 1: Abdomen cutaneous lesions.
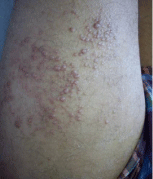
Figure 2: papulo-nodules.
History of the patient revealed that his father and sister were suffering of the same disease (later confirmed as dermatophytic disease caused by Trichophyton rubrum and Trichophyton violaceum on mycological ground). On the other hand, the patient reported previous similar episodes which poorly responded to various drugs including antifungal agents (Fluconazole and itraconazole) with no improvement till now. Later, a genetic study was performed in all members of his family. All the patients with deep dermatophytosis had autosomal recessive CARD9 (caspase recruitment domain– containing protein 9) deficiency [6].
Direct mycological examination of skin and nodular lesions showed 3 to 5μm septated branching hyphae. Culture on Sabouraud dextrose agar of both kinds of samples yielded non-sporulating sterile colonies that couldn’t be identified at the specific level (Figure 3).

Figure 3: Culture Sabouraud dextrose agar, sterile colonies.
In an effort to provide a definitive identification, cultures of the isolates were subjected to PCR targeting the ribosomal internal transcribed space (ITS) domain. Portions of fungal growth were treated for DNA extraction by using QIAamp® DNA Blood Mini Kit 51106 (Qiagen SA, Courtaboeuf, France).
Nucleic acid extraction was performed according to the manufacturers’ instructions. At the end of the procedure, the DNA pellet was dissolved in 50 to 70μl hydration solution. The following oligonucleotides were used as primers:
ITS1: 5’ CAGGCCGGAGGCTGGCCCC 3’
ITS4: 5’ CGAGGACAACCAGAAAAGAT 3’
These primers amplify a 608bp fragment of the ITS regions of rDNA.
PCR was performed on DNA extracts from fungal isolates under the following conditions: the amplification reaction was performed in a total volume of 40μl; the PCR mixture contained 5μl of 10X reaction buffer (Hot star buffer; Qiagen SA, France), 5μl of 2mmol l-1 desoxynucleoside triphosphates containing an equimolar mixture of dATP, dCTP, dGTP and dTTP (Qiagen), 1μl (10 μmol l-1) of each primer, 0.2μl of Hot star Taq polymerase (Qiagen) and 50ng of template DNA. Samples were amplified through 45 cycles in a thermocycler (Thermolyne Amplitron II Series 1091, Barnstead Thermolyne Corporation, Dubuque, IA, USA) as follows: initial denaturation for 15min at 95°C, denaturation for 1mn at 95°C, annealing for 1mn at 54°C and extension for 2mn at 72°C. This was followed by a final extension step for 7min at 72°C.
The resulting amplicons were separated electrophoretically on 2% agarose gel and visualized under UV light after ethidium bromide staining (Figure 4).
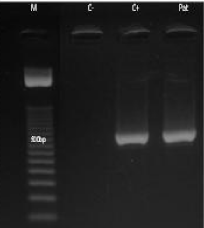
Figure 4: Results of internal transcribed spacer- polymerase chain reaction
from isolate cultures. M: size marker( 50bp DNA ladder); C-:negative control;
C+: positive control; Pat: isolate.
The sequence alignment of the strain showed 100% homology with Trichophyton rubrum registered in Database Trichophyton rubrum strain ATCC MYA-4607 18S.
Discussion
Dermatophytic disease is a rare chronic mycosis characterized by polymorphic cutaneous lesions and possible visceral invasion. It is associated with immunodeficiency, in particular, various inborn errors of interleukin-17 immunity [6] and known to be resistant to antifungals [7]. Correct diagnosis of dermatophytic disease and identification of the causal agent is a major importance as it allows appropriate antifungal treatment to be promptly instituted. Usually, identification of the causing species is based on mycological criteria. The case we report herein shows that identification on the basis of the phenotypical characters may be difficult or impossible when culture yields sterile colonies. In this situation, genome sequencing is needed.
Molecular techniques are much beneficial for fungal identification owing to their rapidity, sensitivity and specificity. Moreover, these methods rely on genetic characters which are more constant than phenotypic ones. In addition, they help to characterize atypical or sterile fungi that are difficult to identify by mycological techniques [8,9,10]. Specific molecular-based techniques for identification of species or even strain in severe fongic infections can be used as possible alternatives for routine identification of fungi [11].
In our study, the identification of the isolate as Trichophyton rubrum could only be achieved after sequencing of ITS region. The pathogenic potential of Trichophyton rubrum is dependent on a variety of local and systemic factors that affect the natural host resistance to dermatophytic infection [12]. In nearly all instances, chronic cutaneous Trichophyton rubrum infections precede deep invasion; tissue invasion is attributed to a presumed alteration of host resistance [13].
The case we report herein highlights the usefulness of molecular techniques in identification of causative agent when the mycological criteria are lacking.
References
- Koga T, Matsuda T, Matsumoto T, Furue M. Therapeutic approaches to subcutaneous mycoses. Am J Clin Dermatol. 2003; 4: 537-543.
- Boudghène-Stambouli O, Mérad-Boudia A. Dermatophytic disease: exuberant hyperkeratosis with cutaneous horns. [Article in French] Ann Dermatol Venereol. 1998; 125: 705-707.
- Souissi A, Ezzine Sebai N, Benmously R, Mokhtar I, Fazaa B et al. Dermatophytic disease in a Tunisian family. Dermatol Ther. 2004; 17: 523-531.
- Cheikhrouhou F, Makni F, Masmoudi A, Sellami A, Turki H, Ayadi A et al. A fatal case of dermatophytic disease with retropharyngeal abscess. Annal Dermatol Vénéréol. 2010; 137: 208-211.
- Kong F, Tong Z, Chen X, Sorrell T, Wang B, Wu Q et al. Rapid identification and differentiation of Trichophyton species, based on sequence polymorphisms of the ribosomal internal transcribed spacer regions, by Rolling-Circle amplification. J .Clin. Microbiol. 2008; 46: 1192-1199.
- Lanternier F, Pathan S, Vincent QB, Liu L, Cypowyj S, Prando C et al. Deep Dermatophytosis and Inherited CARD9 Deficiency. N Engl J Med. 2013; 369: 1704-1714.
- Boudghène-Stambouli O, Mérad-Boudia A. Antifungal agents in dermatophytic disease: failure of griseofulvin, ketoconazole and itraconazole. [Article in French] Bull Soc Pathol Exot. 1990; 83: 170-176.
- Kanbe T. Molecular approches in the diagnosis of dermatophytosis. Mycopathologia. 2009; 18: 801-816.
- Yüksel T, Ilkit M. Identification of rare macroconidia-producing dermatophytic fungi by real-time PCR. Med Mycol. 2012; 50: 346-352.
- Vinod Mootha V, Shahinpoor P, Sutton DA, Xin L, Najafzadeh MJ, de Hoog GS. Identification problems with sterile fungi, illustrated by a keratitis due to a non-sporulating Chaetomium-like species. Med Mycol. 2012; 50: 361-367.
- Kano R, Kusuda M, Nakamura Y, Watanabe S, Tsujimoto H, Hasegawa A. First isolation of Exophiala dermatitidis from a dog: identification by molecular analysis. Veter Microbiol. 2000; 76: 201-205.
- Zhou X, Hu Y, Liu K, Wang L, Wei Q, Han X. Cutaneous and subcutaneous phaeohyphomycosis caused by Exophiala jeanselmei after renal transplantation: a case report. [Article in Chinese]. Nan Fang Yi Ke Da Xue Xue Bao. 2012; 32: 1206-1210.
- Novick NL, Tapia L, Bottone EJ. Invasive Trichophyton rubrum infection in an immunocompromised host. Case report and review of the literature. Am J Med. 1987; 82: 321-325.