
Research Article
Austin J Biomed Eng. 2020; 5(1): 1041.
Study of the Mechanical and Structural Properties of Porcine-Derived Small Intestine Submucosa Decellularized with Triton X-100 at Different Concentrations
Sordelli A1*, Jaunarena JH1, Loresi M1, D’adamo M1, Albite R1, Costa L1, Leonardi L2, Londoño Calderón CL2, Froimowicz P2, DeBadiola F3, Moldes JM3, Giúdice C4, Güeglio G4 and Villoldo GM1,4
1Institute of Translational Medicine and Biomedical Engineering, Hospital Italiano de Buenos Aires, Argentina
2School of Engineering, Institute of Technology in Polymers and Nanotechnology, Argentina
3Pediatric Urology Department, Hospital Italiano de Buenos Aires, Argentina
4Urology Department, Hospital Italiano de Buenos Aires, Argentina
*Corresponding author: Sordelli A, Institute of Translational Medicine and Biomedical Engineering, Hospital Italiano de Buenos Aires, Buenos Aires, Argentina, CONICET, Potosí St 4240, Buenos Aires, C1199ACL, Argentina
Received: September 25, 2020; Accepted: October 20, 2020; Published: October 27, 2020
Abstract
Introduction: Porcine-derived small intestine submucosa has been widely used in tissue engineering. Cell components must be removed to preclude transplant rejection in animal models. Known as decellularization, this type of process can damage collagenous fibers and thus affect the integrity of the extracellular matrix. Purpose: Determine whether structural and physical changes occur in SIS when applying increased concentrations of a non-ionic detergent, Triton X-100, together with fixed concentrations of 0.1% sodium azide. Materials and Methods: SIS was collected, sectioned and placed in distilled water for 24 h before different concentrations of Triton X-100 were applied for 48 h. Samples were removed and stained with hematoxylin and eosin, Masson’s trichrome and Hoechst. Physical characterization was carried out via traction assays; we calculated the rupture strain, ultimate tensile strength and Young’s modulus, and we performed infrared spectroscopy and thermogravimetric analyses. We determined the concentration of remaining DNA.
Results: Regardless of the concentration of detergent used, no cells were revealed at 48 h post-treatment. The patterns of rupture strain, ultimate tensile strength and Young’s modulus, as well as the infrared spectra, exhibited no significant differences between the use of high and low detergent concentrations. A higher detergent concentration results in a lower residual DNA concentration.
Conclusions: It can be concluded that, irrespective of the concentration of Triton X-100 used, at 48 hours post-treatment, decellularization is efficient and does not alter the structural or the physical characteristics of the material. Residual DNA amounts decrease with higher doses and longer treatment times.
Keywords: SIS (Porcine-derived small intestine submucosa); Decellularization; Tissue engineering; Regenerative medicine; Characterization
Abbreviation
SIS: Porcine-derived Small Intestine Submucosa; ECM: Extracellular Matrix; FTIR: Performed Infrared Spectroscopy TGA: Thermogravimetric Analyses; PFA: Paraformaldehyde; H&E: Hematoxylin and Eosin; CEPI: Research Ethics Committee; GAGs: Glucosaninoglycans
Introduction
Matrices derived from natural tissue, such as porcine-derived Small Intestine Submucosa (SIS), have been successfully used both in lab and clinical settings [1-6]. SIS can be easily harvested, processed and handled [2], is biocompatible, degrades easily and resorbs rapidly into the receptor tissue [7,8]. When SIS is transplanted into animals different processes are triggered, such as neovascularization, infiltration with fibroblasts, release of growth factors and other biologically active substances required for restoration processes [9,10].
Natural collagenous matrices need to be decellularized to be used as scaffolds [11]. Several kinds of detergents have been employed at different concentrations for SIS decellularization, but ideal concentrations and exposure times for maximum performance and minimum damage remain unclear. We aimed to analyze the effectiveness of the SIS-decellularization process and find variations in the physical and mechanical properties of SIS under increased concentrations of Triton X-100 combined with fixed concentrations of sodium azide.
Materials and Methods
This study was approved by the Research Ethics Committee of our institution.
SIS collection
SIS was harvested from two male pigs weighing approximately 15kg each. Under aseptic conditions and general anesthesia at 10 minutes post-euthanasia, a median supra-infraumbilical incision was made to obtain a 20cm of jejunum. Mesenteric vessels were bound together at the intestinal margin and the remaining intestine was resealed. The specimen was placed in saline solution and the enteric contents were removed.
SIS processing
Slightly-modified SIS was prepared following Badylack et al.’s protocol [2]. The specimen was placed in saline solution, was cut open at the mesenteric border and the seromuscular layer was removed by careful dissection, moistened gauze was used to mechanically remove the mucosal layer. A 1mm-thick transparent membrane was obtained, sectioned into 5x4 cm rectangles and placed on an orbital shaker (Thermo Scientific) with distilled water for continuous agitation at 200rpm and 37oC for 24 hours. These were then immersed in 250ml screw-cap glass containers (Schott) containing 0.5, 1, 1.5 and 2% Triton X-100 and 0.1% sodium azide solutions and shaken continuously. Samples were collected after 24 and 48 hours.
Histology and morphologic analysis
The matrices were fixed in 4% Paraformaldehyde (PFA), processed and embedded in paraffin. Then they were stored in increased alcohol concentrations (70-100%) and finally cleared with xylene. After this, the paraffin-embedded matrices were sectioned to 4μm thickness.
Routine staining was performed, including Hematoxylin and Eosin (H&E) to validate that decellularization had been satisfactory; Masson’s trichrome to determine the structure of the ECM, and Hoechst to visualize eventually remaining cell nuclei. A Nikon Eclipse E400 microscope was used.
DNA quantification
DNA was extracted from frozen SIS exposed to different detergent concentrations. Each membrane was sectioned into 1g small chunks, which were then placed in a tube. The DNA extraction kit (QIAamp DSP DNA Blood Mini QIAGEN) was filled with a softer tissue lysis buffer (Buffer ATL) and an enzyme (Proteinase K) for SIS molecular disintegration. After incubation at 56 and 37oC respectively, a stronger lysis buffer (Buffer AL) was added to the tube for a 10-minute reincubation at 70oC. Then 100% ethanol was added in a QIAamp Mini spin column of the extraction kit.
After successive spin steps and buffer (AW and AW2) addition, the column was transferred to a tube and DNA was extracted with AE buffer. The DNA in each SIS sample was quantified using the GENESYS 10S spectrophotometer and the ADN.mfx method (which measures wavelengths using a factor) and the result (average after five measurements) was expressed in μg. A curve was plotted with the untreated, water-treated and detergent-treated SIS values, the latter including different detergent concentrations and exposure times.
Hydration time measurement
SIS rectangles were placed on an Explorer Ohaus balance with draft shield to determine their dry weight and were subsequently hydrated in a beaker with saline solution at 25oC. These were then weighed at 5-minute intervals until the matrices’ weight remained unchanged in three consecutive readings and reached a stable weight. The “optimal hydration time” was defined as the first stable reading was obtained.
Mechanical characterization
Uniaxial tensile tests were performed on rectangular decellularized and hydrated SIS samples of 50.8mm in nominal width and varying lengths, i.e. lengths depending on the intestine they had been longitudinally sectioned from. All tests were carried out at room temperature in an Instron 5982 dynamometer with a 1kN load cell at a crosshead speed of 10mm/min until complete fracture. All this data was analyzed using INSTRON’s Bluehill 3® software; stress-strain curves were obtained and calculations were performed to determine the properties of the material. Load-elongation curves were obtained and converted into stress-strain curves using the following: Stress s=F/Ao, where F is the force applied to N and AO is the initial crosssection of each test specimen calculated as its width times its thickness. The thickness was measured using a micrometer (accuracy=0.001 mm) once the test tubes were subjected to a preload of 0.5 N.
Strain ε=Δl/lo, where Δl is the elongation equal to l– lo, and l and lo, the instantaneous and reference length between clamps, respectively.
Young’s modulus was obtained as the initial slope of the stressstrain curve for linear elastic behavior, i.e. where the material deforms but remains recoverable. The ultimate tensile strength (sUTS) was measured as the maximum stress withstood before material fracture, and the strain at break (eb) as the maximum strain experimented by the material. At least six measurements were performed for each group. The average values and their corresponding standard deviations were reported.
SIS lyophilization for Fourier-Transform Infrared (FT-IR) spectroscopy
In order to perform FT-IR spectroscopy and Thermogravimetric Analysis (TGA), samples were first lyophilized in a LABCONCO Freezone 2.5 lyophilizer, model 7670530, for 24-48 hours at room temperature.
FT-IR spectroscopy
Fourier-transform infrared spectra were recorded on a Shimadzu IR Affinity-1 spectrophotometer on net samples. For each spectrum 40 scans were averaged, spectral window was from 450 to 4000 cm-1, at a resolution of 4cm-1. Each sample was measured 6 times in different regions, thus generating a spectrum reflecting the average of all six.
Thermogravimetric analysis
Thermograms for TGA were recorded using a Shimadzu TGA- 50 thermogravimetric analyzer. Sample temperatures were increased once, from room temperature to 600oC. Nitrogen atmosphere (30mL/ min) was used and the heating ramp rate was set at 10oC/min in all cases. Thermal decomposition temperature was determined as the onset on the curve of the first derivative of each thermogram.
Statistics
Data are presented as absolute values or percentages as appropriate. Statistical line, dot and bar charts were used, with 95% confidence intervals represented by the whiskers. A one-way ANOVA was used for group comparison while pair comparison (post hoc) was performed with the Bonferroni correction. For this purpose, IBM SPSS v19.0 (SPSS Inc., Chicago, IL, USA) software was used. Values of p<0.05 were regarded as statistically significant.
Results
SIS collection
At a macroscopic level, a 1 mm-thick, elastic and resistant off white translucent membrane was observed.
Microstructural features
Collagenous fiber structures were preserved after the detergentbased decellularization protocol. Evaluation revealed that decellularization caused variable results (typical of biological samples) and that Triton X-100 treatment causes reduced thickness in some ECM areas. In most cases, however, the structure was preserved, and presented only some regional variations in terms of fiber alignment but no significant differences (i.e., the ultrastructure was preserved). No further damage to the collagenous fibers was observed after increasing the detergent concentration.
Hoechst staining revealed a slight variation in the count of cells attached to SIS (Figure 1). In order to estimate the nuclei count across samples, these were sorted out into groups of n=5 samples per treatment; five points were chosen randomly in each Hoechststained section (with 40x magnification) and averaged. A high nuclei count was observed in the native (untreated) SIS sample, but after placing the SIS in distilled water and applying continuous agitation for 24 hours, a significantly decreased nuclear content was observed although the nuclei count remained high. Histological results revealed a small number or absence of remaining nuclei after the 24-hour protocol (Figure 1C), and a complete absence of nuclear material in the samples following the 48-hour protocol (Figure 1D). The difference observed was statistically significant (p<0.05) when comparing native SIS with each of the groups after 48 hours.
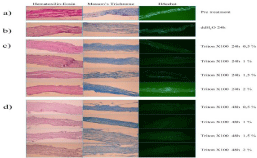
Figure 1: Hoechst staining revealed a slight variation in the count of cells attached to SIS. Histological results revealed a small number or absence of remaining
nuclei after the 24-hour protocol (Figure 1C), and a complete absence of nuclear material in the samples following the 48-hour protocol (Figure 1D).
DNA quantification
DNA quantification was performed on samples extracted from decellularization protocol. A graphic representation is shown in (Figure 2A).
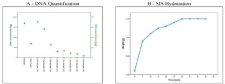
Figure 2: DNA Quantification: DNA quantification was performed on samples extracted from decellularization protocol. A graphic representation is shown in
Figure 2A.
Hydration time measurement: A gradual increase in weight was observed due to water retention, which grew exponentially until the 30-minute time point, when
the weight became stabilized and plateaued out, despite the hydrating procedures still in progress (Figure 2B).
Hydration time measurement
A gradual increase in weight was observed due to water retention, which grew exponentially until the 30-minute time point, when the weight became stabilized and plateaued out, despite the hydrating procedures still in progress (Figure 2B). At t=0 the dry membrane weighed 1.92g; after 5 minutes’ hydration with saline solution increased to 2.08 (+0.16); 10 min reached 2.12g (+0.20); 15 min 2.15g (+0.23); 20 min 2.16g (+0.24); 25 min 2.18 (+0.26); and 30 min, it became stabilized at 2.20g (+0.28). Weight readings were obtained at t=35, 40 and 45 with no variation, thus 30-minute time point as the optimum hydration time.
Uniaxial tensile behavior
Figures 3 A, B and C report values for Young’s modulus (E), which represents the material’s stiffness or response in the elastic range, calculated as the initial slope of the stress-strain curve; the ultimate tensile strength (sUTS), which accounts for the maximum stress withstood by the material; and the strain at break (eb), which represents the material’s ductility or ability to deform, i.e. the maximum strain experienced by the material before fracture. E values ranged from 19 to 34.5MPa, sUTS, from 4 to 9.5MPa and erot, from 55.7 to 118.7%. However, confidence intervals overlapped at some point in all cases, thereby indicating that no significant differences exist among the tensile properties of the different samples.
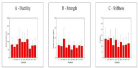
Figure 3: A, B and C report values for Young’s modulus (E), which represents the material’s stiffness or response in the elastic range. No significant differences
exist among the tensile properties of the different samples.
Infrared spectroscopy
Figure 4A shows FT-IR spectra of SIS obtained after different decellularization treatments. All spectra follow similar trends, in terms of the characteristic IR bands, symmetry of signals, shape, and relative intensity. These spectra clearly point to the presence of bands that are commonly associated with SIS, namely Amide I, Amide II and Amide III, as observed between 1660-1672, 1545-1550, and 1239- 1241 cm-1, respectively, in addition to the bands associated with the antisymmetric stretching of CH, between 2930-2949 cm-1.
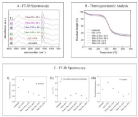
Figure 4: 4A shows FT-IR spectra of SIS obtained after different decellularization treatments. All spectra follow similar trends, in terms of the characteristic IR
bands, symmetry of signals, shape, and relative intensity. Figure 4B shows the TGA results for each SIS sample in the range from 30 to 600oC.
Thermogravimetric analysis
TGA was used to compare the thermal behavior of untreated to that of decellularized SIS sample. Figure 4B shows the TGA results for each SIS sample in the range from 30 to 600oC.
Discussion
The bioactive substances contained in collagenous matrices, such as growth factors, GAGs, cell adhesion molecules and proteins aid cells in the scaffolds to stick to each other, grow and regenerate tissues and organs [12]. Such biocompatibility and regenerative capacity have motivated the use of collagenous matrices, including SIS, in tissue engineering [13,14]. SIS has been safely used for urethra, urinary bladder and tunica albuginea regeneration, among other procedures [2,5].
After the extraction from pig intestine, native intestinal cells must be removed to preclude transplant rejection. Different procedures exist for removing SIS cells, including physical, chemical and physicochemical methods. Physical methods fail to provide adequate decellularization, hence, a chemical method must be added for efficient removal of cells [15]. However, this process can be deleterious to the structure of the collagenous matrix and consequently impact on the scaffold’s effectiveness because of disruption of cell membranes and bonds that bind cells together and to the ECM. Different compounds have been used randomly at different concentrations, but no studies have been made to compare different concentrations and exposure times with the purpose of standardizing the process. We set out to analyze our own decellularization protocol, using 0.1% sodium azide +0.5, 1, 1.5 and 2% concentrations of Triton X-100 for 24 and 48 h with a 24 h distilled water-prewash. The first 24-hour wash with distilled water significantly reduced cell concentrations within matrices, but the addition of Triton/sodium azide solutions enhanced the process to such an extent that 24 hours after applying the detergent few cells were observed, and after 48 hours no nuclei were found. Matrices were invariably decellularized after 48 hours regardless of the detergent concentration employed. Histologically, there were no significant changes across the different groups. We can claim that higher detergent dose represents no extra damage to the matrices, at least as per optical microscopy.
We also explored mechanical properties of SIS. Knowledge of stress and strains in tissues and organs is essential to design devices capable of repairing specific tissues. Manufacturing parameters such as time, dehydration techniques and sterilization methods are also important since they may impact on mechanical properties [15,16]. Our uniaxial tensile test traction assay revealed no significant changes in the parameters analyzed across the samples treated with Triton X-100 at different concentrations. Although a high dispersion was observed in our data set, this may be expected when working with biological matrices. Consequently, we can conclude that the mechanical behavior of the analyzed matrices was not affected by the decellularization protocol.
We used Infrared Spectroscopy (FT-IR) to study the molecular changes and the impact resulting from each decellularization process carried out. There were slight variations in the positioning of the main FT-IR spectrum bands compared to the literature [17-19]. However, these bands were within normal ranges, and the small differences found can only be attributed to the biological nature of the samples. Moreover, in no case did we observe a significant divergence in any of the bands. Furthermore, the fact that Amide II and Amide III bands exhibited similar aspect ratios and intensities, despite having undergone different decellularization treatments, indicates that the triple-helix structure of SIS remained unaffected after those treatments. This result suggests that SIS is stable enough to survive after any of the decellularization treatments described in this study.
TGA is used to characterize biological samples with a high collagenous content and to analyze changes in thermal properties after different chemical treatments [20]. The results suggest a threestep thermal decomposition process of all SIS samples, thus agreeing with previous reports in the literature20. The first process is associated with water loss from the water remaining in the SIS samples even after having carried out drying treatments. The mass loss percentages observed for each sample were attributed to the presence of water still adsorbed and/or absorbed present in a 11.5w/w% even after a 48-hour lyophilization process at room temperature. This indicates that the amount of water in each SIS sample remained almost identical even after having been processed with a decellularization protocol. The second thermal decomposition step, which presents the greatest mass loss, is due to the thermal degradation of the organic components forming the porcine-derived SIS, mainly composed of collagen. The third step is associated with carbonization. The thermal decomposition of all SIS systems was similar, regardless the decellularization treatment carried out. Results suggest that none of the decellularization processes used modified the thermal stability of SIS.
Finally, the study of post-detergent residual DNA amounts is a step toward confirming that a decellularization process is successful. Although DNA amounts decrease dramatically after treatment with distilled water, a greater drop is observed when this is applied for 24 hours and even more when administered for 48 hours (with remaining DNA amounts under 13μg/μL). Therefore, the most efficient method to remove residual DNA is treatment with 2% Triton X-100 for 48 hours.
Conclusion
Decellularization with Triton X-100 is efficient and does not alter structural or physical characteristics of SIS. Regardless of the detergent concentration employed, there were no ultrastructural or histologically significant changes, nor changes in the thermal or tensile properties of any of the groups. Residual DNA amounts decrease with higher doses and longer treatment times: using 2% concentrations for 48 h reduced residual DNA to negligible amounts.
References
- Badylak S. Xenogeneic Extracellular Matrix as a Scaffold for Tissue Reconstruction. Transplant Immunology. 2004; 367-377.
- Badylak SF, Kropp B, McPherson T, Liang H, Snyder PW. Small Intestinal Submucosa: A Rapidly Resorbed Bioscaffold for Augmentation Cystoplasty in a Dog Model. Tissue Engineering. 1998; 379-387.
- Davis NF, Callanan A, McGuire BB, Mooney R, Flood HD, McGloughlin TM. Porcine Extracellular Matrix Scaffolds in Reconstructive Urology: An Ex Vivo Comparative Study of Their Biomechanical Properties. J Mech Behav Biomed Mater. 2011; 4: 375-382.
- Davis NF, Mooney R, Piterina AV, Callanan A, Flood HD, McGloughlin TM. Cell-Seeded Extracellular Matrices for Bladder Reconstruction: An Ex Vivo Comparative Study of Their Biomechanical Properties. The International Journal of Artificial Organs. 2013; 251-258.
- El-Kassaby AW, Retik AB, Yoo JJ, Atala A. Urethral Stricture Repair with an off-the-Shelf Collagen Matrix. J Urol. 2003; 169: 170-173.
- Mantovani F, Tondelli E, Cozzi G, Abed El Rahman D, Spinelli MG, Oliva I, et al. Reconstructive urethroplasty using porcine acellular matrix (SIS): evolution of the grafting technique and results of 10-year experience. Urologia. 2011; 78: 92-97.
- Kanematsu A, Yamamoto S, Noguchi T, Ozeki M, Tabata Y, Ogawa O. Bladder Regeneration by Bladder Acellular Matrix Combined with Sustained Release of Exogenous Growth Factor. J Urol. 2003; 170: 1633-1638.
- Kolker AR, Brown DJ, Redstone JS, Scarpinato VM, Wallack MK. Multilayer Reconstruction of Abdominal Wall Defects with Acellular Dermal Allograft (AlloDerm) and Component Separation. Ann Plast Surg. 2005; 55: 36-41.
- Allman AJ, McPherson TB, Badylak SF, Merrill LC, Kallakury B, Sheehan C, et al. Xenogeneic Extracellular Matrix Grafts Elicit a TH2-Restricted Immune Response. Transplantation. 2001; 71: 1631-1640.
- Zhang J, Wang GY, Xiao YP, Fan LY, Wang Q. The Biomechanical Behavior and Host Response to Porcine-Derived Small Intestine Submucosa, Pericardium and Dermal Matrix Acellular Grafts in a Rat Abdominal Defect Model. Biomaterials. 2011; 7086-7095.
- Gilbert T, Sellaro T, Badylak S. Decellularization of Tissues and Organs. Biomaterials. 2006.
- Chun SY, Lim GJ, Kwon TG, Kwak EK, Kim BW, Atala A, et al. Identification and Characterization of Bioactive Factors in Bladder Submucosa Matrix. Biomaterials. 2007; 28: 4251-4256.
- Hurst RE, Bonner RB. Mapping of the Distribution of Significant Proteins and Proteoglycans in Small Intestinal Submucosa by Fluorescence Microscopy. J Biomater Sci Polym Ed. 2001; 12: 1267-1279.
- Metcalf MH, Savoie FH, Kellum B. Surgical Technique for Xenograft (SIS) Augmentation of Rotator-Cuff Repairs. Operative Techniques in Orthopaedics. 2002; 204-208.
- Syed O, Walters NJ, Day RM, Kim HW, Knowles JC. Evaluation of Decellularization Protocols for Production of Tubular Small Intestine Submucosa Scaffolds for Use in Oesophageal Tissue Engineering. Acta Biomater. 2014; 10: 5043-5054.
- Liao J, Joyce EM, Sacks MS. Effects of Decellularization on the Mechanical and Structural Properties of the Porcine Aortic Valve Leaflet. Biomaterials. 2008, 29: 1065-1074.
- Ghassemi T, Saghatoleslami N, Mahdavi-Shahri N, Matin MM, Gheshlaghi R, Moradi A. A Comparison Study of Different Decellularization Treatments on Bovine Articular Cartilage. J Tissue Eng Regen Med. 2019.
- Poornejad N, Schaumann LB, Buckmiller EM, Momtahan N, Gassman JR, Ma HH, et al. The Impact of Decellularization Agents on Renal Tissue Extracellular Matrix. J Biomater Appl. 2016, 31; 521-533.
- Deeken CR, Bachman SL, Ramshaw BJ, Grant SA. Characterization of Bionanocomposite Scaffolds Comprised of Mercaptoethylamine- Functionalized Gold Nanoparticles Crosslinked to Acellular Porcine Tissue. J Mater Sci Mater Med. 2012, 23; 537-546.
- Wang Q, Cao C, Liao J, Jiang J, Wang L, Li X, et al. Development of Three-Dimensional Porous Architecture and Biocompatibility Based Poly (p-Dioxanone) Composite with Small Intestinal Submucosa for Soft Tissue Repair and Reconstruction. Journal of Biomaterials and Tissue Engineering. 2017; 1225-1234.