
Research Article
Austin J Pharmacol Ther. 2021; 9(3).1140.
Identification of KUP/HAK/KT Potassium Transporter Gene Family in Wild Carrot (Daucus carota) and Its Phylogenetic Analysis
Mazhar MW*, Batool MS, Saleem T, Mahmood J, Saif S, Waqas N, Tahirv H and Mazhar F
Department of Bioinformatics and Biotechnology, Government Collage University, Pakistan
*Corresponding author: Muhammad Waqar Mazhar, Department of Bioinformatics and Biotechnology, Government Collage University, 38000 Faisalabad, Pakistan
Received: May 11, 2021; Accepted: June 04, 2021; Published: June 11, 2021
Abstract
Potassium is essential macronutrient of plant. It is uptaken by plant through channels and transporters. Plants specie genomes contains a number of KUP/ HAK/KT transporters having the primary function to mediate K+ fluxes. In this study,we have identified 13 gene members of KUP/HAK/KT transporter gene family. In most of the plant species, these genes have been characterized but uncharacterized in D. carota. This study has been done to identify KUP/HAK/KT gene family in D. carota plant species to study its phylogeny. This gene family is important for potassium uptake and play an important role in translocation, osmotic potential regulation, plant development and growth. Different tools like MEGA 7.0.21, pfam, SMART and NCBI-BLASTp has been used to characterize the gene family. This study covers the phylogeny and evolution of KUP/HAK/KT transporters in D. carota with reference to A. thaliana.
Keywords: K+ ion; A. thaliana; D. carota; KUP/HAK/KT; Phylogeny
Introduction
Potassium ion (K+) being the essential macronutrient is found very important for various aspects of plant life. In plants, potassium concentration may reach up to 8% of dry cell weight. Potassium is the most plentiful cation that maintains the electrical charge balance and transport of sugar and nitrate in plant cells [1]. It may act as protein synthesis stabilizer or enzyme activator. It is involved in metabolic and physiological processes, such as photosynthesis, cellular osmoregulation and respiration [2]. potassium deficiency highly affects the plant growth and development as they pertain many abiotic stresses like salth stress, drought stress and cold stress [3]. Potassium is found in bulk amount in earth crust and the uptake of potassium from soil is not as good as it to be because potassium is not in ionic form. The concentration of potassium at root surface is mostly lower than in bulk soil solution which may fall down to μM range. To enhance the potassium uptake, plant have developed various mechanism for potassium acquisition with >1000-fold concentration gradients [4].
Usually there are two mechanisms for potassium uptake in plants. These are high affinity transporter system via potassium transporters and low affinity potassium transporters via potassium channels. On the basis of function and structure, potassium transporters are divided into five families: 1) shaker channels; 2) TDK (tandem porek+; 3) HAK (high affinity K+) /KUP (K+ uptake permease) /KT (K+ transporter); 4) HKT transporters; 5) CPAs (cation protonantiporters). KUP/HAK/KT is the largest and widely distributed family in bacteria, fungi, and plants. The homology with bacterial KUP and fungal HAK transporters show that the plants KUP/HAK/ KT transporter members AtKUP1 and HvHAK1 were first cloned from Arabidopsis and Barley. Comparative genomic analysis showed that 13, 27 and 27 KUP/HAK/KT genes were identified in Arabidopsis, Rice and Maize respectively [5]. Mutation analysis has shown that the 8th transmembrane domain and C-terminus of KUP/HAK/KT gene family has a key role in determining the K+ transport capacity. A series of KUP/HAK/KT genes were identified in past decades and a number of physiological roles of these genes were characterized in plant species. At present, KUP/HAK/KT genes have been identified in other species like poplar (Populus trichocarpa), tomato (Solanum lycopersicum), pear (Pyrus bretschneideri) and soybean (Glycine max) [6] (Table 1).
Regulation of cell size, auxin distribution or osmotic stress adaptation
[29]
Mediate K+ fluxes
[25]
Subcellular localization (plasma membrane, tonoplast, or other endomembranes).
[26,27[
Expression patterns (root meristems, vascular tissues, guard cells, fruits, or specialized organs such as flytraps).
[26,28]
Table 1: Some major functions relted to KUP/HAK/KT.
Expression of KUP/HAK/KT is regulated by K+ starvation, abiotic stresses and phytohormones including ABA, cytokinin, ethylene and NAA. These KUP/HAK/KT proteins shows a great variety of subcellular localization in tonoplast, plasma membrane and endoplasmic reticulum. This gene family have role in plant growth and development. The ubiquitous nature of these genes show that they are very important for plants as they help them in survival in potassium poor environment and in nutrients uptake [4].
Daucus carota is a popular taproot vegetable, a plant rich in many bioactive pigments like carotenoids and lutein, cultivated all over the world [7]. Carotenoid found in flowers and fruits of plant playing major role in photosystems of higher plants. It can enhance the pollination ability of plant and seed dispersal activity. It accumulates in carrot taproot being the major nutrient of carrot taproot and also contributes to carrot taproot color [8].
The analysis is done in comparison with the features of this gene family in Arabidopsis. This comprehensive study may provide important information about lineage of KUP/HAK/KT gene family in Daucus carota to other species.
Materials and Methods
Identification and evolutionary analysis of KUP/HAK/KT genes in D. carota
The KUP/HAK/KT potassium transporting genes of D. carota were retrieved from NCBI database (https://www.ncbi.nlm.nih.gov/). The protein sequence of KUP/HAK/KT potassium transporting genes were retrieved from already known protein sequence of potassium transporter genes in A. thaliana. Protein blast (Blastp) was performed against genome database and GenBank to retrieve the protein and DNA sequences of D. carota. A. thaliana gene sequence were used as query sequence against non-redundant protein sequence of D. carota. The retrieved data was manually checked to eradiacte the non identical or false positives. The sequences having similarity less than 60% were discarded. Both candidate protein sequences were examined for conserved domains using pfam domain (http://pfam.xfam.org/) [9,10] and SMART (SMART: Main page (embl-heidelberg.de) [11] and sequences were filtered to get finer collection of nonredundant gene sequences present in D. carota. This process was done to eliminate the sequences having no conserved domains. Multiple sequence alignment was done to confirm the conserved domains in the target plant species D. carota. The full length KUP/HAK/KT protein from A. thaliana and D. carota were aligned in MEGA 7.0.21 software and neighbor joining evolutionary tree were constructed with bootstrapping value of 1000 for phylogenetic analysis [12,13]. NCBI database (https://www.ncbi.nlm.nih.gov) was used to recover all genomic information of these potassium transporting genes like exon numbers, chromosomal location of gene, proteina and RNA accession numbers, amino acid length and ORF length.
Gene structure analysis
To predict gene structure and their schematic representation, an online freely available server called Gene Structure Display server (GSDS) was used (http://gsds.gao-lab.org/) [14].
Trace the different restriction maps and multiple sequence alignment
Different restriction maps were generated of each sequence by using SerialCloner2-6 software [15]. It generated different graphic maps of each sequence of identifies gene sequence. For multiple sequence alignment, a web-based tool called ClustalW (https://www. genome.jp/tools-bin/clustalw) was used to align the sequence of target specie [16].
Results and Discussions
Identification of KUP/HAK/KT in D. carota and conserved domain profile
The KUP/HAK/KT protein sequences of A. thaliana, C. arietinum and O. sativa (Reference species) were retrieved from NCBI. The retrieved AtKUP/HAK/KT sequences were used in NCBI-BLAST search against D. carota sequence. After making all queries, 13 members of KUP/HAK/KT gene family were identified out of more than 100 sequences and named as 1) DcKUPHAKKT2, 2) DcKUPHAKKT3, 3) DcKUPHAKKT5.1, 4) DcKUPHAKKT5.2, 5) DcKUPHAKKT6.1 6) DcKUPHAKKT6.2, 7) DcKUPHAKKT6.3, 8) DcKUPHAKKT7.1, 9) DcKUPHAKKT7.1, 10) DcKUPHAKKT8, 11) DcKUPHAKKT10, 12) DcKUPHAKKT11 and 13) DcKUPHAKKT12. The nomenclature was proposed according to functions of homologous Arabidopsis genes. The evolution process tells that the similar structure and similar function of genes arise through replication event and gene duplication is the process which leads towards evolution [17]. The analysis revealed that D. carota contains 13 members of KUP/ HAK/KT gene family as in A. thaliana. The AtHAK2, AtHAK6 and AtHAK11 are responsible for salinity response because their expression gets affected when salt concentration is increased [1]. Salinity is caused by reduced Na+ controlled by AtHKT1 and AtHAK5 which is regulated by external K+ concentration [18].
The gene location on the chromosome from top to bottom was analysed. From the analysis, it was analyzed that all of the genes are present on chromosomes having number of 1-9 except 5 and 8. No gene is present on chromosome number 5 and 8. Chromosome location is helpful to find the location of gene on chromosome. Similarly, the exon count analysis reveals that its average range is 8-11, four genes have 9 exons, 4 genes have 8 exons, two genes contain 10 exons and 1 gene contains 11 exons. The presence of K_trans domain in the identified sequence were confirmed using an online tool pfam (http://pfam.xfam.org/) [10] and SMART (SMART: Main page (embl-heidelberg.de) [19].
Pfam and SMART server searches revealed that the most conserved domain in all sequences is K_trans which show that it belongs to potassium transporter gene family (Table 1 and Table 2).
Proposed names
Gene locus
Protein accession#
RNA accession#
Exons
Chr#
ORF length
Amino acid length
Start of genomic location
Conserved domains in protein sequence
DcKUPHAKKT2
DCAR_019813
KZM92822.1
XM_017399321.1
9
6
2349
782
36563422
K_trans
DcKUPHAKKT3
DCAR_014456
XP_017247390.1
XM_017391901.1
9
4
2349
782
21753814
K_trans
DcKUPHAKKT1
DCAR_007762
XP_017231954.1
XM_017376465.1
8
2
2277
758
36900730
K_trans
DcKUPHAKKT2
DCAR_023264
XP_017255168.1
XM_017399679.1
9
6
2475
824
196829
K_trans
DcKUPHAKKT1
DCAR_007096
XP_017232096.1
XM_017376607.1
8
2
2403
800
31460152
K_trans
DcKUPHAKKT2
DCAR_031093
XP_017224518.1
XM_017369029.1
9
9
2340
779
32656696
K_trans
DcKUPHAKKT3
DCAR_024139
XP_017218890.1
XM_017363401.1
8
7
2343
780
9796649
K_trans
DcKUPHAKKT1
DCAR_001786
XP_017225142.1
XM_017369653.1
10
1
2535
844
20502453
K_trans
DcKUPHAKKT2
DCAR_021857
XP_017258440.1
XM_017402951.1
10
6
2532
843
19110565
K_trans
DcKUPHAKKT8
DCAR_011216
XP_017241396.1
XM_017385907.1
8
3
2325
774
33669288
K_trans
DcKUPHAKKT0
DCAR_001295
XP_017229797.1
XM_017374308.1
8
1
2364
787
4149046
K_trans
DcKUPHAKKT1
DCAR_007784
XP_017236764.1
XM_017381275.1
11
2
2352
783
37055812
K_trans
DcKUPHAKKT12
DCAR_007095
XP_017235837.1
XM_017380348.1
9
2
2526
841
31449412
K_trans
Table 2: Whole Genomic Information about identified genes.
Phylogenetic analysis
To characterize the evolutionary relationship between KUP/ HAK/KT gene family of A. thaliana, C. arietinum, O. sativa and D. carota, MEGA 7.0.21 was used to generate the rooted tree by the neighbor joining method by the alignment of KUP/HAK/KT from both species (Figure 1). It is concluded that the KUP/HAK/KT gene family is common in almost all plant species. The evolutionary tree showed a close relationship between O. sativa, C. arietinum, A. thaliana and D. carota. This gene family helps in mediating the inward transport of K+ from Dilute K+ solutions as shown in OsHAK1 and OsHAK5 transporters [20]. As the KUP/HAK/KT transporters are multi gene family. For example, Arasbidopsis has 13 KUP/HAK/ KT genes and Oryza sativa has 27 KUP/HAK/KT genes in total [21]. Investigation shows that KUP/HAK/KT can be divided into four clusters which is confirmed by the phylogenetic analysis of full-length sequences of KUP/HAK/KT transporter gene family. The most studied gene members of KUP/HAK/KT transporter gene family belongs to clusterI and clusterII [22]. ClusterI gene members play essential role in potassium uptake when potassium concentration is low. So, it may be designated as high affinity potassium uptake transporters in Arabidopsis which may be have the same role in D. carota too [4]. Phylogenetic analysis results indicate that evolutionary branches were constant with their assigned gene names. Introns have been found essential traits of eukaryotic gene families during the evolutionary process of multiple gene families [23].
Figure 1: Genome wide analysis conducted in plants so far.
Gene structure analysis
Usually it is found that the position of introns/exons in orthologous genes are commonly well conserved in evolutionary times, but the structure of introns/exons is comparatively less conserved in paralogus genes. To study the structural differences of KUP/HAK/KT gene family we used Gene Structure Display tool (http://gsds.gaolab.org/) which analyzed the arrangements of introns and exons by comparing CDS sequences and Genomic sequences [24-29]. It shows that DcKUPHAKKT5.2 has more exonic regions than others. In contrast the longest intronic regions are present in DcKUPHAKKT10 and DcKUPHAKKT7.2 gene. The highest exon count is 11 in DcKUPHAKKT11 (Figure 2). It has been reported that AtKUP2 gene plays important role in maintianing cell size. Because K+ is the major solute in cell so any diminishing effect to the potassium homeostasis leads to the weakend cell turgor which causes reduction in cell size. Similarly, the gene in D. carota will also affect the cell size [4] (Figure 3).
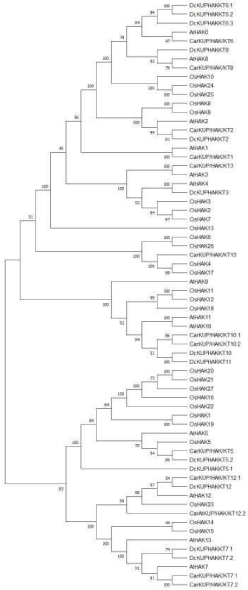
Figure 2: Phylogenetic analysis of A. thaliana, O. sativa and D. carota.
The phylogenetic tree of potassium transporter genes was constructed with
MEGA 7 using Neighbor-Joining method with 1,000 bootstrap replicates.

Figure 3:
Mapping Restriction sites
To find the restriction sites of identified sequence, SerialCloner2-6 was used. The genomic sequence of all the 13 gene members were saved in FASTA format and then each sequence was submitted to SerilCloner2-6 software and a graphic map of restriction sites were generated (Figure 4A-D).

Figure 4A: Restrion map performed by SerialCloner software 2-6.

Figure 4B:
Figure 4C:

Figure 4D:
Conclusion
Potassium is the macronutrient of plants, it is uptaken by plants through various mechanism. High affinity potassium uptake through Potassium transporter is one of them. This study reported 13 gene members of KUP/HAK/KT potassium transporter gene family in D. carota (wild carrot). This study covers the genomic information, domain prediction, gene structure prediction and phylogenetic analysis. These genes showed a close phylogenetic relationship with A. thaliana, C.arietinum and O. sativa. This gene family is important for potassium transport. Different software like NCBIBLASTp, pfam, SMART and SerialCloner2-6 was used. Gene structure display server software were also used to conduct gene structure analysis. MEGA 7.0.21 software was used to make the phylogenetic tree for evolutionary study.
References
- Li W, Xu G, Alli A, Yu L. Plant HAK/KUP/KT K+ transporters: Function and regulation. Seminars in Cell & Developmental Biology. 2018; 74: 133–141.
- Liang M, Gao Y, Mao T, Zhang X, Zhang S, Zhang H, et al. Characterization and Expression of KT/HAK/KUP Transporter Family Genes in Willow under Potassium Deficiency, Drought, and Salt Stresses. BioMed Research Interational. 2020: e2690760.
- Xie Q, Ma L, Tan P, Deng W, Huang C, Liu D, et al. Multiple High-Affinity K+ Transporters and ABC Transporters Involved in K+ Uptake/Transport in the Potassium-Hyperaccumulator Plant Phytolacca acinosa Roxb. Plants. 2020; 9: 470.
- Grabov A. Plant KT/KUP/HAK Potassium Transporters: Single Family – Multiple Functions. Annals of Botany. 2007; 99: 1035–1041.
- Feng X, Wang Y, Zhang N, Wu Z, Zeng Q, Wu J, Wu X, et al. Genomewide systematic characterization of the HAK/KUP/KT gene family and its expression profile during plant growth and in response to low-K+ stress in Saccharum. BMC Plant Biology. 2020; 20: 20.
- MaoNi C, QingYu W, JinYu Z, ZhiYong Z, Jie D, YaXin Y. Identification and expression analysis of KUP/HAK/KT potassium transporter gene family in soybean [Glycine max (L.) Merr.]. Acta Botanica Boreali-Occidentalia Sinica. 2017; 37: 239–249.
- Fikselová M, Šilhár S, Marecek J, Francáková H. Extraction of carrot (Daucus carota L.) carotenes under different conditions. Czech Journal of Food Sciences. 2008; 26: 268–274.
- Ma J, Xu Z, Tan G, Wang F, Xiong A. Distinct transcription profile of genes involved in carotenoid biosynthesis among six different color carrot (Daucus carota L.) cultivars. Acta Biochimica et Biophysica Sinica. 2017; 49: 817–826.
- Bateman A, Birney E, Cerruti L, Durbin R, Etwiller L, Eddy SR, et al. The Pfam Protein Families Database. Nucleic Acids Research. 2002; 30: 276–280.
- El-Gebali S, Mistry J, Bateman A, Eddy SR, Luciani A, Potter SC, et al. The Pfam protein families database in 2019. Nucleic Acids Research. 2019; 47: D427–D432.
- Letunic I, Doerks T, Bork P. SMART 7: Recent updates to the protein domain annotation resource. Nucleic Acids Research. 2012; 40: D302–D305.
- Hall BG. Building Phylogenetic Trees from Molecular Data with MEGA. Molecular Biology and Evolution. 2013; 30: 1229–1235.
- Kumar S, Nei M, Dudley J, Tamura K. MEGA: A biologist-centric software for evolutionary analysis of DNA and protein sequences. Briefings in Bioinformatics. 2008; 9: 299–306.
- Hu B, Jin J, Guo A-Y, Zhang H, Luo J, Gao G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics. 2015; 31: 1296–1297.
- Amara A. a Globin Hemoglobinopathy as A Case Study: Mutants and Prototype Mutants: Are They Absent a0, Reduced a+ or Recombinant? Cellular & Molecular Medicine. 2017.
- Thompson JD, Gibson TJ, Higgins DG. Multiple Sequence Alignment Using ClustalW and ClustalX. Current Protocols in Bioinformatics. 2003; 231-232.
- Shen C, Yuan J. Genome-Wide Investigation and Expression Analysis of K+- Transport-Related Gene Families in Chinese Cabbage (Brassica rapa ssp. Pekinensis). Biochemical Genetics. 2021; 59: 256–282.
- Wang Q, Guan C, Wang P, Lv M-L, Ma Q, Wu G-Q, et al. AtHKT1;1 and AtHAK5 mediate low-affinity Na+ uptake in Arabidopsis thaliana under mild salt stress. Plant Growth Regulation. 2015; 75: 615–623.
- Schultz J, Copley RR, Doerks T, Ponting CP, Bork P. SMART: A web-based tool for the study of genetically mobile domains. Nucleic Acids Research. 2000; 28: 231–234.
- Santa-María GE, Oliferuk S, Moriconi JI. KT-HAK-KUP transporters in major terrestrial photosynthetic organisms: A twenty years tale. Journal of Plant Physiology. 2018; 226: 77–90.
- Gierth M, Mäser P, Schroeder JI. The Potassium Transporter AtHAK5 Functions in K+ Deprivation-Induced High-Affinity K+ Uptake and AKT1 K+ Channel Contribution to K+ Uptake Kinetics in Arabidopsis Roots. Plant Physiology. 2005; 137: 1105–1114.
- Gupta M, Qiu X, Wang L, Xie W, Zhang C, Xiong L, et al. KT/HAK/KUP potassium transporters gene family and their whole-life cycle expression profile in rice (Oryza sativa). Molecular Genetics and Genomics: MGG. 2008; 280: 437–452.
- Liang M, Gao Y, Mao T, Zhang X, Zhang S, Zhang H, et al. Characterization and expression of KT/HAK/KUP transporter family genes in willow under potassium deficiency, drought, and salt stresses. BioMed Research International. 2020.
- Aslam M, Fakher B, Jakada BH, Zhao L, Cao S, Cheng Y, et al. Genome- Wide Identification and Expression Profiling of CBL-CIPK Gene Family in Pineapple (Ananas comosus) and the Role of AcCBL1 in Abiotic and Biotic Stress Response. Biomolecules. 2019; 9: 293.
- Ahn SJ, Shin R, Schachtman DP. Expression of KT/KUP Genes in Arabidopsis and the Role of Root Hairs in K+ Uptake. Plant Physiology. 2004; 134: 1135–1145.
- Osakabe Y, Arinaga N, Umezawa T, Katsura S, Nagamachi K, Tanaka H, et al. Osmotic Stress Responses and Plant Growth Controlled by Potassium Transporters in Arabidopsis. The Plant Cell. 2013; 25: 609–624.
- Rigas S, Debrosses G, Haralampidis K, Vicente-Agullo F, Feldmann KA, Grabov A, et al. TRH1 Encodes a Potassium Transporter Required for Tip Growth in Arabidopsis Root Hairs. The Plant Cell. 2001; 13: 139–151.
- Scherzer S, Böhm J, Krol E, Shabala L, Kreuzer I, Larisch C, et al. Calcium sensor kinase activates potassium uptake systems in gland cells of Venus flytraps. Proceedings of the National Academy of Sciences. 2015; 112: 7309– 7314.
- Véry A-A, Nieves-Cordones M, Daly M, Khan I, Fizames C, Sentenac H. Molecular biology of K+ transport across the plant cell membrane: What do we learn from comparison between plant species? Journal of Plant Physiology. 2004; 171: 748–769.