Mini Review
Finding Epigenetic Determinants of the Metabolic Syndrome
Zhang Y1*, Cerjak D1, Ali O2
1Department of Medicine, Medical College of Wisconsin, USA
2Department of Pediatrics, Medical College of Wisconsin, USA
*Corresponding author: Zhang Y, Department of Medicine, Medical College of Wisconsin, USA,
Received: October 03, 2014; Accepted:November 21, 2014; Published: November 27, 2014
Citation: Zhang Y, Cerjak D, Ali O. Finding Epigenetic Determinants of the Metabolic Syndrome. Austin J Endocrinol Diabetes. 2014;1(6): 1029. ISSN: 2381-9200.
Abstract
Metabolic Syndrome (MetS) substantially increases one’s risk for type 2 diabetes (T2D) and cardiovascular disease. It now affects more than one third of adults in the U.S. and has similar impact on other societies globally. Results from recent genome-wide association studies (GWAS) have suggested that the inherited variance in anindividuals’risk of expressing MetS traits cannot be completely explained by variation in the primary sequence of the genome; mechanisms beyond the genetic sequence variants are increasingly compelling for researchers in the field. Epigenetic modifications such as DNA methylation and histone modifications are hypothesized to play important roles in the pathophysiology of diseases including MetS and may explain some of the missing heritability. Recent pilot studies conducted in humans and animals have also suggested epigenetic changes such as CpG methylation modify one’s susceptibility to developing MetS in response to prenatal and postnatal environmental exposures. Although these findings are intriguing, more work is needed in order to unravel a map of epigenetic determinants of MetS.
Keywords: Metabolic Syndrome; DNA Methylation; Epigenetic; Obesity; Environmental Cues
Introduction
Chronic diseases such as cancer, type 2 diabetes (T2D), metabolic syndrome (MetS), cardiovascular disease and dementia constitute the most common health problems seen in developed societies ( increasingly, in developing societies) and their prevalence increases with age in all populations [1-4]. It is well established that environmental exposures, especially in early life, can alter the risk of various chronic diseases later in life [5, 6] and while the mechanisms involved in this “programming” of future risk are not yet understood in detail, epigenetic changes are believed to play an important role in this process [7, 8]. Epigenetic mechanisms are also postulated to be involved in modifying the risk of MetS secondary to postnatal exposures and may explain the “missing heritability” of chronic diseases like MetS. In this mini-review, we discuss the historical context of the concept of MetS epigenetic, the recent evidence and our current opinions about this rising field.
The Metabolic Syndrome (MetS) is a form of obesity characterized by a cluster of phenotypes that includes increased abdominal fat mass, impaired insulin responsiveness, dyslipidemia with increased plasma triglycerides and decreased HDL-cholesterol, increased blood pressure and elevated circulating cytokines and adipokines [9]. It is estimated to affect 34% of adult Americans [10] and adds an extra $2,000 per person in annual health care costs [11]. Its prevalence is low in childhood and increases with advancing age [12, 13].
The prevalence of MetS-associated cardiovascular (CV) risk factors is relatively low during early childhood but increases during adolescence and thereafter tends to persist into adulthood [12-15]. We have observed a similar trend in adolescents in our cross-sectional study population [16]. It is also known that the atherosclerotic process starts in childhood and is accelerated in individuals who are insulin resistant, dyslipidemic and/or show signs of systemic inflammation [17-19]. At a molecular level, the mechanisms by which obesity leads to the development of insulin resistance, dyslipidemia and associated phenotypes are poorly understood but necessarily involve long-term changes in genetic regulation and gene expression. Since the underlying DNA sequence remains unchanged, these changes in gene regulation and function must be mediated by epigenetic mechanisms. These mechanisms, including methylation of CpG sites of DNA, are some of the most important processes by which genetic function is regulated and altered by development and by the external environment [20-26].
Why study epigenetic in MetS
Epigenetic mechanisms, which involve DNA and histone modifications, mediate the interaction between gene and environment throughout the lifespan; while the underlying genetic sequence does not change, environmental influences can alter epigenetic marks and thus alter gene expression and induce long term changes in phenotype and disease susceptibility [27]. The gradual accumulation of epigenetic changes in critical genes may contribute to the observed age-related increase in the prevalence of various chronic disorders [28-31]. Epigenetic changes are known to be heritable across more than one generation of offspring in plants and mammals [32-37] and there is evidence that transgenerational epigenetic inheritance also occurs in humans [38-41]. Such transgenerational inheritance of epigenetic states may contribute to the observed inherited risk of various chronic disorders, including metabolic disorders [42].
DNA methylation is one of the most extensively studied epigenetic mechanisms and plays an important role in the process of development and differentiation [43]. There is evidence from both human and animal sources that prenatal nutritional deprivation can permanently alter DNA methylation at multiple loci and these changes play a role in the observed alteration of future risk of chronic diseases like obesity, insulin resistance and diabetes [44-50]. It is also known that DNA methylation patterns continue to change after birth, at least partly in response to environmental influences [51- 53]. Environmental factors can alter epigenetic features and change the future behavior of target cells and may therefore play a role in susceptibility to chronic diseases, including MetS [54-56]. In the next section, we will briefly review the role epigenetic mechanisms play in the development of MetS.
Epigenetic, prenatal exposures and MetS
Several studies have shown persistent epigenetic changes in humans who face nutritional stress during prenatal life and early childhood [45, 57-58]. Environmental exposures in early life can influence MetS phenotypes later in life through epigenetic mechanisms. Children exposed to prenatal famine and low birth weight has increased risk of T2D, hypertension and other CV disease [50, 58]. Furthermore, a large and extensive epidemiological study of humans who were prenatally exposed to famine during the Dutch Hunger Winter in 1944–1945 showed that 60 years later they had less DNA methylation of the imprinted IGF2 gene compared with their unexposed, same-sex siblings [44].
Transgenerational epigenetic influences on MetS
Recent genome-wide association studies (GWAS) of MetS and its traits individually [59-62] revealed tens to hundreds of DNA variants that are significant but with small effect size in explaining the total heritable variance observed in each phenotype [59, 61]. Increasing evidence now shows that environment-induced genetic effects can pass transgenerationally without changes occurring in the primary DNA sequence [63] and this epigenetic trait can be transmitted up to the fifth generation [64]. Therefore, some of the familial risk of MetS may actually be epigenetic in origin. In rats, female offspring of overweight fathers (on a high fat diet) had an early onset of impaired insulin secretion and glucose tolerance that worsened with time compared to controls [42]. A recent human study showed paternal pre-conceptual obesity was associated with hypomethylation of IGF2 in newborns [40].
Global epigenetics markers are shown to be inherited from one generation to the next. In a family study conducted by McRae et al., by using subjects of 117 families the authors show the average heritability of DNA methylation measured at CpG sites with no known SNPs is estimated to be 0.187 [41]. Carless et al. conducted another study in Mexican American families with high prevalence of obesity and T2D and found 24% of CpG sites tested had nominal evidence of heritabilityand the average level of heritability of these sites is 36% [65]. Some of these heritable CpGs reside within genes of known functions in metabolism [Carless M.A., personal communication]. In our family cohort of Northern European descent, we have also observed a significant portion of the epigenome is heritable, including genes known to play roles in obesity, T2D and MetS [Zhang et al., unpublished].
Postnatal transient and long-lasting epigenetic changes associated with MetS
It is also known that DNA methylation patterns continue to change after birth, at least partly in response to environmental influences [51-53]. For example, studies show that identical twins have broadly similar epigenetic profiles in-utero but these profiles gradually diverge as they get older [20, 66, 67]. Female subjects exposed to the Dutch Famine Winter when they are in young exhibited a 1.3 to 1.6 fold increased risk of type 2 diabetes as compared to unexposed women [68]. Several studies have looked at the effect of aging on genome-wide DNA methylation in adults and these studies show that age-dependent methylation changes are found in a variety of tissues and correlate well enough with age that the methylation status of selected loci can be used to predict the age of a subject [53, 69-71]. Our data show that within families at high risks for developing obesity-related metabolic disorders, there are age-associated genomic loci densely situated near genes that function in the hedgehog signaling and the maturity-onset diabetes of the young pathways (MODY) [in review]. This suggests a novel mechanism underlying the gradual deleterious effects of multiple genes and their interactions with nutrition over time, which may contribute to obesity and its complications. Our study sheds light on the relationship between ageing and increased prevalence of obesity, T2D and their related abnormalities and a dynamic epigenetic landscape that changes throughout the life span.
Exercise is an environmental factor that can also influence both DNA methylation and CV disease and obesity risk. A recent study on exercise epigenetics shows that DNA methylation of genes in retinol metabolism, calcium signaling pathways and with known functions in muscle biology and T2D decreased after exercise [72]. Some of these exercise-associated methylation changes accompanied differential gene expressions [72]. In another study using adipose tissues, 18 obesity and 21genes exhibited differential methylation at CpG loci in response to exercise.
These authors suggest exercise induces genome-wide changes in DNA methylation in human adipose tissue, potentially affecting adipocyte metabolism [73].
Conclusion
MetS has reached epidemic proportion in the last three decadesand is still on the risein essentially all populations. Epigenetic mechanism such as genomic CpG methylation may play an important role in individual differences expressing MetS traits as these epigenetic markers are able to integrate environmental cues into gene expression.
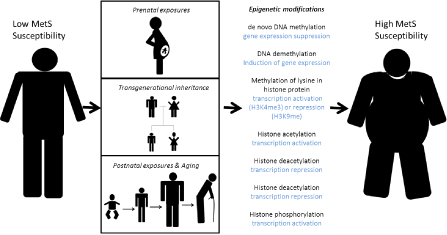
Although it is a novel field, we see increasing amounts of interesting data supporting this theory (Figure 1).
Future Directions
More genome-wide searches for MetS relevant epigenetic variants using human populations that are well-characterized for MetS are needed to map to the interesting regions for follow-up studies. Studies that combine human sample with animal models will be instrumental in delineating the mechanisms where by identified candidates work. In both humans and animals, it will be essential to know the epigenetic states of both surrogate tissues and MetS targets such as adipose tissue, liver and muscle in relation to MetS expression. In addition, population studies using longitudinal samples as well as ones focusing on the effects of environmental cues such as diet and lifestyles will help us find novel targets for risk evaluation, diagnosis and treatment in the clinic.
References
- Lozano R, Naghavi M, Foreman K, Lim S, Shibuya K, Aboyans V, et al. Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet. 2012; 380: 2095-2128.
- Guariguata L, Whiting DR, Hambleton I, Beagley J, Linnenkamp U, Shaw JE . Global estimates of diabetes prevalence for 2013 and projections for 2035. Diabetes Res Clin Pract. 2014; 103: 137-149.
- Ward BW, Schiller JS . Prevalence of multiple chronic conditions among US adults: estimates from the National Health Interview Survey, 2010. Prev Chronic Dis. 2013; 10: E65.
- Hanson M, Gluckman P . Developmental origins of noncommunicable disease: population and public health implications. Am J Clin Nutr. 2011; 94: 1754S-1758S.
- Santos MS, Joles JA . Early determinants of cardiovascular disease. Best Pract Res Clin Endocrinol Metab. 2012; 26: 581-597.
- Kelishadi R, Poursafa P . A review on the genetic, environmental, and lifestyle aspects of the early-life origins of cardiovascular disease. Curr Probl Pediatr Adolesc Health Care. 2014; 44: 54-72.
- Heerwagen MJ, Miller MR, Barbour LA, Friedman JE . Maternal obesity and fetal metabolic programming: a fertile epigenetic soil. Am J Physiol Regul Integr Comp Physiol. 2010; 299: R711-722.
- Suter MA, Ma J, Vuguin PM, Hartil K, Fiallo A, Harris RA, Charron MJ4 . In utero exposure to a maternal high-fat diet alters the epigenetic histone code in a murine model. Am J Obstet Gynecol. 2014; 210: 463.
- Grundy SM, Brewer, Jr HB, Cleeman JI, Smith Jr SC, Lenfant C. Definition of Metabolic Syndrome Report of the National Heart, Lung, and Blood Institute/American Heart Association Conference on Scientific Issues Related to Definition. Circulation.2004; 109: 433-438.
- Ervin RB . Prevalence of metabolic syndrome among adults 20 years of age and over, by sex, age, race and ethnicity, and body mass index: United States, 2003-2006. Natl Health Stat Report. 2009; : 1-7.
- Boudreau DM, Malone DC, Raebel MA, Fishman PA, Nichols GA, Feldstein AC, Boscoe AN . Health care utilization and costs by metabolic syndrome risk factors. Metab Syndr Relat Disord. 2009; 7: 305-314.
- Steinberger J, Moran A, Hong CP, Jacobs DR Jr, Sinaiko AR . Adiposity in childhood predicts obesity and insulin resistance in young adulthood. J Pediatr. 2001; 138: 469-473.
- Juhola J, Magnussen CG, Viikari JS, Kähönen M, Hutri-Kähönen N, Jula A, et al . Tracking of serum lipid levels, blood pressure, and body mass index from childhood to adulthood: the Cardiovascular Risk in Young Finns Study. J Pediatr. 2011; 159: 584-590.
- Park MH, Falconer C, Viner RM, Kinra S . The impact of childhood obesity on morbidity and mortality in adulthood: a systematic review. Obes Rev. 2012; 13: 985-1000.
- Brufani C, Ciampalini P, Grossi A, Fiori R, Fintini D, Tozzi A,et al . Glucose tolerance status in 510 children and adolescents attending an obesity clinic in Central Italy. Pediatr Diabetes. 2010; 11: 47-54.
- Ali O, Cerjak D, Kent JW Jr, James R, Blangero J, Zhang Y . Obesity, central adiposity and cardiometabolic risk factors in children and adolescents: a family-based study. Pediatr Obes. 2014; 9: e58-62.
- McGill HC Jr, McMahan CA, Herderick EE, Malcom GT, Tracy RE, Strong JP . Origin of atherosclerosis in childhood and adolescence. Am J Clin Nutr. 2000; 72: 1307S-1315S.
- Gast KB, Tjeerdema N, Stijnen T, Smit JW, Dekkers OM . Insulin resistance and risk of incident cardiovascular events in adults without diabetes: meta-analysis. PLoS One. 2012; 7: e52036.
- Miller M . Dyslipidemia and cardiovascular risk: the importance of early prevention. QJM. 2009; 102: 657-667.
- Dolinoy DC, Huang D, Jirtle RL . Maternal nutrient supplementation counteracts bisphenol A-induced DNA hypomethylation in early development. Proc Natl Acad Sci U S A. 2007; 104: 13056-13061.
- Anderson OS, Nahar MS, Faulk C, Jones TR, Liao C, Kannan K, et al. Epigenetic responses following maternal dietary exposure to physiologically relevant levels of bisphenol A. Environ Mol Mutagen. 2012; 53: 334-342.
- Fraga MF, Ballestar E, Paz MF, Ropero S, Setien F, Ballestar ML, et al . Epigenetic differences arise during the lifetime of monozygotic twins. Proc Natl Acad Sci U S A. 2005; 102: 10604-10609.
- Oates NA, van Vliet J, Duffy DL, Kroes HY, Martin NG, Boomsma DI, et al . Increased DNA methylation at the AXIN1 gene in a monozygotic twin from a pair discordant for a caudal duplication anomaly. Am J Hum Genet. 2006; 79: 155-162.
- Huang TH, Perry MR, Laux DE . Methylation profiling of CpG islands in human breast cancer cells. Hum Mol Genet. 1999; 8: 459-470.
- Faulk C, Dolinoy DC . Timing is everything: the when and how of environmentally induced changes in the epigenome of animals. Epigenetics. 2011; 6: 791-797.
- Dolinoy DC, Jirtle RL . Environmental epigenomics in human health and disease. Environ Mol Mutagen. 2008; 49: 4-8.
- Delcuve GP, Rastegar M, Davie JR . Epigenetic control. J Cell Physiol. 2009; 219: 243-250.
- Wang G, Walker SO, Hong X, Bartell TR, Wang X . Epigenetics and early life origins of chronic noncommunicable diseases. J Adolesc Health. 2013; 52: S14-21.
- Tammen SA, Friso S, Choi SW . Epigenetics: the link between nature and nurture. Mol Aspects Med. 2013; 34: 753-764.
- Barros SP, Offenbacher S . Epigenetics: connecting environment and genotype to phenotype and disease. J Dent Res. 2009; 88: 400-408.
- van Otterdijk SD, Mathers JC, Strathdee G . Do age-related changes in DNA methylation play a role in the development of age-related diseases? Biochem Soc Trans. 2013; 41: 803-807.
- Hauser MT, Aufsatz W, Jonak C, Luschnig C . Transgenerational epigenetic inheritance in plants. Biochim Biophys Acta. 2011; 1809: 459-468.
- Morgan HD, Sutherland HG, Martin DI, Whitelaw E . Epigenetic inheritance at the agouti locus in the mouse. Nat Genet. 1999; 23: 314-318.
- Cropley JE, Suter CM, Beckman KB, Martin DI . Germ-line epigenetic modification of the murine A vy allele by nutritional supplementation. Proc Natl Acad Sci U S A. 2006; 103: 17308-17312.
- Waterland RA, Travisano M, Tahiliani KG . Diet-induced hypermethylation at agouti viable yellow is not inherited transgenerationally through the female. FASEB J. 2007; 21: 3380-3385.
- Waterland RA, Travisano M, Tahiliani KG, Rached MT, Mirza S . Methyl donor supplementation prevents transgenerational amplification of obesity. Int J Obes (Lond). 2008; 32: 1373-1379.
- Csaba G, Karabélyos C, Inczefi-GondaA, Pállinger E. Three-generation Investigation on Serotonin Content in Rat Immune Cells long after ß-endorphin Exposure in Late Pregnancy. HormMetab Res. 2005; 37: 172-177.
- Heard E, Martienssen RA . Transgenerational epigenetic inheritance: myths and mechanisms. Cell. 2014; 157: 95-109.
- HocherB. More than genes: the advanced fetal programming hypothesis. J Reprod Immunol. 2014; pii: S0165-0378(14)00030-8. doi: 10.1016/j.jri.2014.03.001
- Soubry A, Hoyo C, Jirtle RL, Murphy SK . A paternal environmental legacy: evidence for epigenetic inheritance through the male germ line. Bioessays. 2014; 36: 359-371.
- McRae AF, Powell JE, Henders AK, Bowdler L, Hemani G, Shah S, et al . Contribution of genetic variation to transgenerational inheritance of DNA methylation. Genome Biol. 2014; 15: R73.
- Ng SF, Lin RC, Laybutt DR, Barres R, Owens JA, Morris MJ . Chronic high-fat diet in fathers programs β-cell dysfunction in female rat offspring. Nature. 2010; 467: 963-966.
- Smith ZD, Meissner A . DNA methylation: roles in mammalian development. Nat Rev Genet. 2013; 14: 204-220.
- Heijmans BT, Tobi EW, Stein AD, Putter H, Blauw GJ, Susser ES, et al. Persistent epigenetic differences associated with prenatal exposure to famine in humans. Proc Natl Acad Sci U S A. 2008; 105: 17046-17049.
- Tobi EW, Lumey LH, Talens RP, Kremer D, Putter H, Stein AD, et al . DNA methylation differences after exposure to prenatal famine are common and timing- and sex-specific. Hum Mol Genet. 2009; 18: 4046-4053.
- Reynolds RM, Jacobsen GH, Drake AJ . What is the evidence in humans that DNA methylation changes link events in utero and later life disease? Clin Endocrinol (Oxf). 2013; 78: 814-822.
- Szyf M, Bick J . DNA methylation: a mechanism for embedding early life experiences in the genome. Child Dev. 2013; 84: 49-57.
- Gomes MV, Pelosi GG . Epigenetic vulnerability and the environmental influence on health. Exp Biol Med (Maywood). 2013; 238: 859-865.
- Drake AJ, McPherson RC, Godfrey KM, Cooper C, Lillycrop KA, Hanson MA, et al. An unbalanced maternal diet in pregnancy associates with offspring epigenetic changes in genes controlling glucocorticoid action and foetal growth. Clin Endocrinol (Oxf). 2012; 77: 808-815.
- Godfrey KM, Sheppard A, Gluckman PD, Lillycrop KA, Burdge GC, McLean C, et al. Epigenetic gene promoter methylation at birth is associated with child's later adiposity. Diabetes. 2011; 60: 1528-1534.
- Ziller MJ, Gu H, Müller F, Donaghey J, Tsai LT, Kohlbacher O, et al . Charting a dynamic DNA methylation landscape of the human genome. Nature. 2013; 500: 477-481.
- Bell JT, Tsai PC, Yang TP, Pidsley R, Nisbet J, Glass D, et al . Epigenome-wide scans identify differentially methylated regions for age and age-related phenotypes in a healthy ageing population. PLoS Genet. 2012; 8: e1002629.
- Alisch RS, Barwick BG, Chopra P, Myrick LK, Satten GA, Conneely KN, et al. Age-associated DNA methylation in pediatric populations. Genome Res. 2012; 22: 623-632.
- Gapp K, von Ziegler L, Tweedie-Cullen RY, Mansuy IM. Early life epigenetic programming and transmission of stress-induced traits in mammals: How and when can environmental factors influence traits and their transgenerational inheritance? Bioessays. 2014; 36: 491-502.
- Portha B, Fournier A, Kioon MD, Mezger V, Movassat J . Early environmental factors, alteration of epigenetic marks and metabolic disease susceptibility. Biochimie. 2014; 97: 1-15.
- Sun H, Kennedy PJ, Nestler EJ . Epigenetics of the depressed brain: role of histone acetylation and methylation. Neuropsychopharmacology. 2013; 38: 124-137.
- Cruickshank MN, Oshlack A, Theda C, Davis PG, Martino D, Sheehan P, et al. Analysis of epigenetic changes in survivors of preterm birth reveals the effect of gestational age and evidence for a long term legacy. Genome Med. 2013; 5: 96.
- Wehkalampi K1, Muurinen M, Wirta SB, Hannula-Jouppi K, Hovi P, Järvenpää AL, Eriksson JG . Altered Methylation of IGF2 Locus 20 Years after Preterm Birth at Very Low Birth Weight. PLoS One. 2013; 8: e67379.
- Speliotes EK, Willer CJ, Berndt SI, Monda KL, Thorleifsson G, Jackson AU, et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat Genet. 2010; 42: 937-948.
- Heard-Costa NL, Zillikens MC, Monda KL, Johansson A, Harris TB, Fu M, et al . NRXN3 is a novel locus for waist circumference: a genome-wide association study from the CHARGE Consortium. PLoS Genet. 2009; 5: e1000539.
- Kraja AT, Vaidya D, Pankow JS, Goodarzi MO, Assimes TL, Kullo IJ, et al. A bivariate genome-wide approach to metabolic syndrome: STAMPEED consortium. Diabetes. 2011; 60: 1329-1339.
- Sladek R, Rocheleau G, Rung J, Dina C, Shen L, Serre D, et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature. 2007; 445: 881-885.
- Nadeau JH . Transgenerational genetic effects on phenotypic variation and disease risk. Hum Mol Genet. 2009; 18: R202-210.
- Anway MD, Cupp AS, Uzumcu M, Skinner MK. Epigenetic transgenerational actions of endocrine disruptors and male fertility. Science.2005; 308, 1466-1469.
- Carless MA, Kulkarni H, Kos MZ, Charlesworth J, Peralta JM, Göring HH, et al. Genetic effects on DNA methylation and its potential relevance for obesity in Mexican Americans. PLoS One. 2013; 8: e73950.
- Talens RP, Christensen K, Putter H, Willemsen G, Christiansen L, Kremer D, et al . Epigenetic variation during the adult lifespan: cross-sectional and longitudinal data on monozygotic twin pairs. Aging Cell. 2012; 11: 694-703.
- Wong CC, Caspi A, Williams B, Craig IW, Houts R, Ambler A, et al . A longitudinal study of epigenetic variation in twins. Epigenetics. 2010; 5: 516-526.
- van Abeelen AF, Elias SG, Bossuyt PM, Grobbee DE, van der Schouw YT, Roseboom TJ, et al . Famine exposure in the young and the risk of type 2 diabetes in adulthood. Diabetes. 2012; 61: 2255-2260.
- Hannum G, Guinney J, Zhao L, Zhang L, Hughes G, Sadda S, et al . Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol Cell. 2013; 49: 359-367.
- Day K, Waite LL, Thalacker-Mercer A, West A, Bamman MM, Brooks JD, et al . Differential DNA methylation with age displays both common and dynamic features across human tissues that are influenced by CpG landscape. Genome Biol. 2013; 14: R102.
- Teschendorff AE, Menon U, Gentry-Maharaj A, Ramus SJ, Weisenberger DJ, Shen H, et al. Age-dependent DNA methylation of genes that are suppressed in stem cells is a hallmark of cancer. Genome Res. 2010; 20: 440-446.
- Nitert MD, Dayeh T, Volkov P, Elgzyri T, Hall E, Nilsson E, et al . Impact of an exercise intervention on DNA methylation in skeletal muscle from first-degree relatives of patients with type 2 diabetes. Diabetes. 2012; 61: 3322-3332.
- Rönn T, Volkov P, Davegårdh C, Dayeh T, Hall E, Olsson AH, et al . A six months exercise intervention influences the genome-wide DNA methylation pattern in human adipose tissue. PLoS Genet. 2013; 9: e1003572.
Epigenetic mechanisms underlying MetS etiology. Current literature supports three scenarios where epigenetic modifications may have affected an individual’s susceptibility to develop MetS traits through prenatal exposures, transgenerational inheritance and postnatal exposures and aging.