
Special Article - Cell Death and Autophagy
Austin J Mol & Cell Biol. 2015; 2(1): 1004.
Intersection of Apoptosis and Autophagy Cell Death Pathways
Masayuki Noguchi¹*, Noriyuki Hirata¹, Tatsuma Edamura¹, Satoko Ishigaki¹ and Futoshi Suizu¹
¹Division of Cancer Biology, Institute for Genetic Medicine, Hokkaido University, Japan
*Corresponding author: Masayuki Noguchi, Division of Cancer Biology, Institute for Genetic Medicine, Hokkaido University, N15, W7, Kita-ku, Sapporo 060- 0815, Japan,
Received: April 24, 2015; Accepted: May 20, 2015; Published: May 22, 2015
Abstract
The balance between cell survival and death is a critical parameter in the regulation of cell and tissue homeostasis. Autophagy is an evolutionarily conserved mechanism for the gross disposal and recycling of intracellular proteins in mammalian cells. Autophagy also kills cells under certain conditions, in a process called autophagic cell death; this involves pathways and mediators different from those of apoptosis. Therefore, three different mechanisms of cell death have been identified in mammalian cells; namely, apoptosis (type I), autophagic cell death (type II), and necrosis (type III). Whether and how these different processes of cell death interconnected each other has not been fully clarified. In this review we discuss the evidence supporting a mechanistic link especially focusing between apoptosis and autophagy associated cell death—including the possibility of cross–talk between the relevant signaling pathways—that could serve to maintain cellular homeostasis in mammals.
Keywords: Apoptosis; Autophagy; Autophagy-related genes (ATG); Cell Death
Introduction
In recent decades, insight into the molecular regulation of autophagy in mammalian cells has come from the discovery and functional analysis of Autophagy-Related Gene (ATG). Autophagy is an evolutionally conserved homeostatic process for intracellular degradation by which intracellular proteins are sequestered in a double–membrane–bound autophagosome and delivered to the lysosome during stress conditions; this process facilitates both degradation and recycling of intracellular proteins in mammalian cells. The molecular machinery of autophagy co-ordinates diverse aspects of cellular and organismal responses to other dangerous stimuli such as infection [1,2]. Defective autophagy underlies a wide variety of human disease and physiology including cancer, neurodegeneration, and infectious diseases [3-5]. Mammalian orthologues of ATG family proteins have been identified and various functions of ATG proteins have been elucidated, including how these proteins control the formation of autophagosomes. Although autophagy was originally characterized as a cytoprotective process in yeast under starvation conditions, it is now thought to be a form of cell death [6,7] along with the two classical mechanisms of apoptosis and necrosis in mammalian cells.
Three possible mechanisms for cell death have been known to exist in mammalian cells, namely apoptosis (type I cell death), autophagic cell death (type II cell death), and necrosis (type III cell death). Apoptotic cell death (type I cell death) is characterized by rounding up of the cell and reduction of cell volume, chromatin condensation, nuclear fragmentation, no modification of cytoplasmic organelle, and plasma membrane blebbing without involvement of gene activity. Since autophagy is thought to be a pro-survival pathway, whether or not autophagy indeed induce cell death is still under debate. However, under certain circumstances, autophagy can induce cell death (type II cell death) which is characterized by presence of massive autophagic vacuole in the cytoplasm. Necrosis (type III cell death) is most classical form of cell death with characteristic morphological feature of a gain of cell volume, swelling of organelles with plasma membrane rupture without blebbing.
There is accumulating evidence for cross-talk in the regulation of apoptosis and induction of autophagy [8-12]. The present review examines how these three types of cell death interact in mammalian cells.
Three types of cell death in mammalian cells
Three different mechanisms of cell death are known to exist in mammalian cells, namely apoptosis (type I), autophagy (type II), and necrosis (type III) [6,7,13] (Table 1). Apoptosis, a form of programmed cell death [14], was originally distinguished from traumatic or necrotic cell death based on cytological features by electron microscopy. Research over the past two decades has elucidated the major molecules in apoptotic signaling pathways from the plasma membrane to the nucleus; it is known to be triggered by ligands such as Tumor Necrosis Factor (TNF) and Tnf-Related Apoptosis-Inducing Ligand (TRAIL) that activate cell death receptors such as Fas-Associated Protein with Death Domain (FADD) [15].
Types of Cell Death
Characteristic
features
Apoptosis
(Type I)
Autophagy
(Type II)
Necrosis
(Type III)
Nucleus
Reduction of volume,
Chromatin condensation,
Nuclear fragmentation
Absence of chromatin condensation
Swelling, Chromatin fragmented
Cytoplasm
Little/no modification of cytoplasmic organelles
Presence of apoptotic body
Presence of massive autophagic vacuole
Swelling of organelles, Subsequent loss of content
Mitochondria
Morphologically normal initially
Possibly involved
with autophagic molecule
Morphologically aberrant
Plasma Membrane
Blebbing
-
Disrupted cell membrane
Cell volume
Decreased
-
Increased
Caspases Activation
Involved
No involvement
No involvement
Gene activation
Required
In some cases
No involvement
Lysosome
unaffected
Active executer
Abnormal
Inflammation
No
Possibly
Marked
Three types of cell death are known to be present in mammalian cells. Although three types of cell death are not necessarily classified by their morphological character, these three types of cell death have distinct morphologically distinct features to some extent [6,7,13].
Morphologic features of Apoptotic cell death (type I cell death) is characterized by rounding up of the cell and reduction of cell volume, chromatin condensation, nuclear fragmentation, no modification of cytoplasmic organelle, plasma membrane blebbing. In addition functionally apoptotic cell death does not involve lysosome degradation, but requires specific gene activity.
In contrast to apoptotic cell death autophagic cell death (type II cell death) is characterized not necessarily by its morphological features. Since autophagy is thought to be a pro-survival pathway, whether or not autophagy indeed induce cell death is still under debate. However, it is generally accepted that in certain circumstances, autophagy occasionally induce cell death which can be “Death with autophagy”. Autophagic cell death is characterized by presence of massive autophagic vacuole in the cytoplasm, by the absence of chromatin condensation and lacking of caspase activation.
Necrosis (type III cell death) is the most classical form of cell death. Necrosis is occasionally associated with impairment of blood supply and therefore massive cellular infarction, hence subsequent loss of intracellular content by lysis. Characteristic morphological feature of necrosis includes a gain of cell volume, swelling of organelles with plasma membrane rupture, but no signs of blebbing. In striking contrast to apoptosis, no specific gene activity is required in necrosis. It is of note that necrosis is also known to be taken place as a consequence of apoptosis [13].
Table 1: Three types of cell death in mammalian cells.
Autophagy is a conserved mechanism that functions in the degradation and recycling of proteins in mammalian cells [3-5]. It was originally characterized as a process by which cells recycle cytoplasmic contents and defective organelles during cellular stress conditions such as nutrient starvation in yeast. In autophagy (which refers to macroautophagy, in contrast to microautophagy and chaperone-mediated autophagy), cytosolic proteins and organelles such as mitochondria are sequestered within double-membrane vesicles to facilitate the formation of autophagosomes that fuse with the lysosome. ATG products function cooperatively in this process at multiple steps. To date, more than 30 mammalian orthologues of ATG family proteins have been identified, and their functions have been characterized primarily by gene targeting technology. Autophagy is essential for maintaining cellular homeostasis, and mutations in ATG or dysregulation of autophagy pathways underlie various pathological conditions [3-5].
Cell death associated with autophagy has been proposed in mammalian cells [6,11,16,17]. However, one fundamental question is how and whether excessive bulk digestion which is occurring at the lysosome can necessarily cause cell death [18].The molecular events associated with apoptosis—including caspase activation, chromatin condensation, DNA cleavage, or plasma membrane degeneration— is well known [15]. Increasing lines of evidence indicate that these molecular mechanisms may be recruited by an alternative, caspaseindependent form of programmed cell death, named autophagic type II cell death [6,11,14,16,17]. Following growth factor withdrawal, Bax-/-Bak-/- cells activate autophagy, undergo progressive atrophy, and ultimately succumb to death. The observation supported the idea that growth factor signal transduction is required to direct the utilization of sufficient exogenous nutrients to maintain cell viability [11]. However, the molecular processes that occur between the lysosomal degradation of cytosolic components leading autophagic cell death are poorly characterized [19]. Moreover, it remains to be confirmed whether autophagy induces cell death in a physiological setting [10].
Necrosis is a cell death mechanism that does not require specific death receptor signaling [20]. It is often triggered by massive occlusion of the blood supply such as in cerebral or myocardial infarction, which leads to widespread but subsequent loss of intracellular content by lysis. Characteristic morphological changes in necrotic cells include a gain of cell volume, swelling of organelles with plasma membrane rupture without blebbing. It is also recognized that necrosis is considered as to be no gene activity are required. But necrosis is known to be taken place as a consequence of apoptosis [13]. Although necrosis is considered to be a passive process, in certain situations necroptosis can actively induce programmed cell death [21,22]. I need to emphasize that it is unclear whether any mechanistic interactions are occurring between the autophagic and necrotic pathways in mammalian cells.
Cross-talk between apoptosis and autophagy pathways
Many recent studies have focused on potential cross-talk between the three cell death pathways [6,11,16,17,23] (Figure 1). In particular, interactions between autophagy and the other two mechanisms (apoptosis and necrosis) have been focused its attention in recent years [9,21,22]. Inhibition of macroautophagy is shown to trigger apoptosis [24]. Although autophagy is cytoprotective effect on starvation condition by lysosomal degradation of intracellular component, autophagy is normally a cell-survival pathway involving the degradation and recycling of obsolete, damaged, or harmful macromolecular assemblies. However in some experimental settings, autophagy is believed to induce, or more precisely, autophagy is associated with cell death, so called type II cell death [25,26].
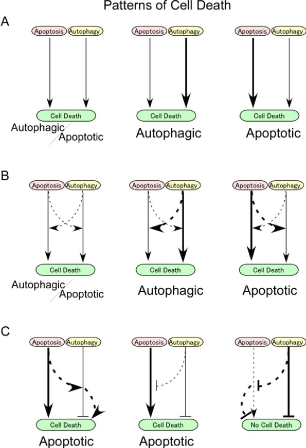
Figure 1: Relationship between apoptotic and autophagy related cell death
in mammalian cells.
Three types of cell death are generally believed to be present in mammalian
cells namely apoptosis, autophagic cell death and necrosis. In particular,
interactions between autophagy and apoptosis have been focused its
attention in recent years [9,21,22]. Autophagy may enhance cell death
caused by apoptosis; alternatively, it may induce cell death independently of
apoptosis or necrosis. Whether and how these types of induction cross talk
each other in physiological settings in vivo remain to be solved.
Assuming both autophagy and apoptosis (and/or necrotic) induce cell death,
it is possible that these mechanisms of cell death operate independently of
each other (A). In this scenario, depending on the strength of the cell deathinducing
signal, the phenotype of cell death can be apoptotic or autophagic;
however, if they occur simultaneously then the manifestation of each
mechanism will be less obvious. In this situation, autophagy interacts with
apoptotic pathways to coordinately induce cell death. The second possibility
is that autophagy and apoptotic signals cooperatively functions to induce
cell death (B). Under this circumstance, if the relative strength of death
inducing signal of autophagy is weaker than the cell death inducing signal by
apoptosis, the cell will exhibit a phenotype that is either purely apoptotic or
mixed apoptotic and autophagic. The third possibility is that even if autophagy
has cytoprotective roles, depending on the relative strength of induction of
apoptosis for death execution vs. autophagy for cell survivals, the phenotype
can be either autophagic, apoptotic or a combination of them depending on
the relative strength of cell death induced by apoptosis or cytoprotective
effect by autophagy (C).
If autophagy has solely cytoprotective function, and induction of apoptosis and induction of autophagy are taking place simultaneously, there are two possible phenotypic manifestations can be appeared depending on the relative strength of autophagy for cytoprotective vsinduction of cell death through apoptosis. Depending on the relative strength of autophagy vs apoptosis, for example, if the autophagy could not fully prevent apoptosis-induced cell death, the possible apparent manifestation of cell death can be autophagic cell death or possibly apoptosis based on the relative dominance of distinct morphological changes.
Cells undergoing autophagic cell death do not exhibit chromatin condensation as in apoptotic cells, but show massive vacuolization of the cytoplasm and accumulation of double-membrane autophagosomes, with little or no uptake by phagocytic cells; these features allow the two processes to be distinguished morphologically [6,7,13].
However, an open question is whether autophagy is cell survival mechanism or indeed functioning as an actively killing mechanism in mammalian cells [10,27,28].
Technical limitations that preclude the unambiguous identification of morphological features present a challenge in the study of autophagic cell death [29-32]. In experimental systems, autophagic cells are typically identified by visualizing microtubuleassociated protein 1 Light Chain (LC) 3 puncta by microscopy, or by assessing the levels of LC3 II/I (which reflects the phosphatidylserine conversion of LC3 at the lysosomal membrane [32,33]) or p62 (which is a marker of autophagic degradation) by western blotting [34,35]. However, these approaches are based solely on the process of lysosomal degradation during autophagy; markers for other steps in the pathway would be useful, for instance those that are specific to the process of cell death induced by autophagy. In contrast, methods for evaluating the various steps of apoptosis are well established, including annexin V staining to detect plasma membrane damage, the detection of cleaved caspase levels to assess caspase activation, and terminal deoxynucleotided UTP nick end labeling to examine DNA damage. Thus, technical limitations remain a hindrance in the clear distinction between autophagic and apoptotic cell death [29,31,32].
Autophagy was originally described in yeast cells as a mechanism for cell survival [18,36] that can counter cell death, which implies an interaction between autophagic, apoptotic, and necrotic signaling pathways. In yeast, autophagy is solely a mechanism for cell survival through amino acid recycling [37]. It is recently revealed that apoptosis in yeast is functioning to induce cell death [38]. In contrast, autophagy in mammalian cells, which has been characterized in past decades, appears to be much more complex. In mammalian cells, primary function of autophagy is thought to be cell survival mechanism. However, autophagy can induce cell death or alternatively, mammalian cells are dying associated with autophagy in certain conditions [6,11,16,17,25,26].
Autophagy may enhance cell death caused by apoptosis; alternatively, it may induce cell death independently of apoptosis or necrosis. In contrast to autophagy, the defined molecular regulation for cell death cascade from the death receptor and its downstream signal is well characterized in the apoptotic pathways —starting from the activation of the death receptor, which is followed by a downstream signaling cascade including the involvement of mitochondria, subsequent caspase activation, and DNA cleavage [39]. However, pro-apoptotic signals such as TRAIL [40], TNF [41], and FADD [42] are also known to induce autophagy. Pro-apoptotic signals, which is promoting or causing apoptosis, participate in a cascade that lead to culminate in cleavage of a set of proteins, resulting in disassembly of the cell for apoptosis [43]. Moreover, UVRAG human homolog of yeast Vps38, has been shown to inhibit BAX to regulate apoptosis [44]. Ectopic expression of Beclin-1(ATG6) suppresses cell death while reduction of Beclin-1 levels by siRNA sensitizes cells to TRAILinduced cell death [45].
If it is assumed that autophagy in mammalian cells functions primarily to promote cell survival (cytoprotective), then depending on the relative strength of cell death-inducing signals acting on the cell, the processes of apoptosis or necrosis may prevail over the protective effects by autophagy and the fate of the cell could be interpreted as incomplete apoptosis with an autophagic phenotype; however, these processes may nonetheless kill the cell [10,46].
Mitochondria as a point of intersection for autophagic and apoptotic cell death
Mitochondria are shown to be playing an important role in the induction of apoptosis through the cytochrome c release via the disruption of mitochondrial outer membrane potential [47]. Mitochondrial Transmembrane Potential (MTP) plays a key role in the regulation of apoptotic cell death machinery and that AKT regulates this process [6,48-50].
Accumulating evidences have shown that mitochondria as an intersection of autophagy and apoptosis (Figure 2). Mitochondriaassociated proteins may also be responsible for interactions between the autophagic and apoptotic pathways [8]. Calpain mediated cleavage of ATG5 can modulate the Bcl-2/Beclin1 protein complex, a key regulator for apoptosis at mitochondria [51]. Bcl-2 and Beclin1 physically interacted each other at the mitochondrial outer membrane [52]. The lack of ATG12-ATG3 complex formation produces an expansion in mitochondrial mass and inhibits cell death mediated by mitochondrial pathways [53].
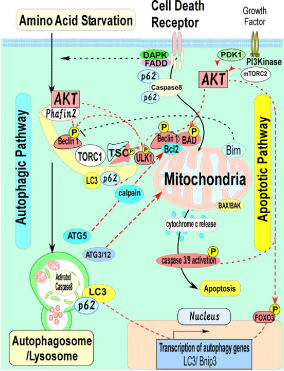
Figure 2: Autophagic and apoptotic pathways converge in mitochondria.
Mitochondria would provide an ideal molecular platform of counter regulation
of autophagic cell death vs. apoptotic cell death. In this regard, mitochondriaassociated
proteins may also be responsible for interactions between the
autophagic and apoptotic pathways [8]. Numbers of ATG families such as
ATG5, ATG3/12 have influence on mitochondrial function. Calpain mediated
cleavage of ATG5 can modulate the Bcl-2/Beclin1 protein complex, a key
regulator for apoptosis at mitochondria [51]. Bcl-2 and Beclin1 physically
interact each other at the mitochondrial outer membrane [52]. The lack of
ATG12-ATG3 complex formation produces an expansion in mitochondrial
mass and inhibits cell death mediated by mitochondrial pathways [53].
The autophagy protein ATG12 associates with anti-apoptotic Bcl-2 family
members to promote mitochondrial-mediated apoptosis [64]. Anti-apoptotic
protein Bcl-2 is shown to inhibit Beclin 1-dependent autophagy [56]. Beclin1
(ATG6) and ULK1 (ATG1) are both involved in autophagosome formation and
are direct substrates of AKT [63,68]. Fork head transcription factor FoxO3,
a substrate of AKT, is shown to activate autophagy related genes including
LC3 and Bnip3 [79]. Moreover, Beclin1 or Bim, which influence on autophagic
process are shown to be localized at the mitochondrial membrane. More
precisely, these molecules are shown to be involved in the regulation of
vesicle formation of stage of autophagosomal formation. Further, mTORC1,
a crucial autophagy regulator, is shown to directly phosphorylate ULK1
(ATG1) and ATG13 [75]. AKT and TSC (tuberous sclerosis complex) are
also localized at lysosome [61,71]. These observations are consistent that
induction of autophagy and apoptosis can be cross talked at the intersection
of mitochondrial and vesicle nucleation stage of autophagosome formation of
autophagy at ER-mitochondria contact site [83].
DAP-kinase (death associated protein kinase) is a mediator of endoplasmic reticulum stress-induced caspase activation and simultaneously involved in the regulation of autophagic cell death [54]. Bim, another member of anti-apoptotic Bcl-2 family proteins colocalized at mitochondria and also shown to inhibit autophagy via Beclin1 [55-57]. Consistent with the notion that phosphorylation of autophagy–related proteins is an additional aspect of autophagy regulation [58], phosphorylation of Beclin1 on T119 by DAP-kinase also reduces the Bcl-2-Beclin1 interaction and activates autophagy [57].
At the outer membrane of mitochondria AKT can phosphorylate BAD [59], which then release activated forms of Bcl-2 to prevent the subsequent cytochrome c release for downstream caspase activation [60]. Involvement of AKT in the regulation of autophagy was suggested by the fact that AKT directly phosphorylate wide varieties of autophagy and apoptotic regulatory molecules localized at either mitochondria or autophagosome including ULK1 (Unc-51 like autophagy activating kinase 1, ATG1), ATG6 (Beclin1), BAD, TSC2 (Tuberous sclerosis complex 2) [23,61]. Cross talk between apoptosis and autophagy by caspase-mediated cleavage of Beclin1, a substrate of AKT [62,63]. The autophagy protein ATG12 associates with anti-apoptotic Bcl-2 family members to promote mitochondrialmediated apoptosis [64]. Alternatively, activated caspase8, which is associated with p62, is known to be proteolyzed via lysosome [65]. Further, lysosomal membrane permeabilization induces cell death in a mitochondrion-dependent fashion [66]. It is noteworthy that mitochondria itself also targeted by lysosomal degradation called mitophagy [67].
It is possible that AKT and its phosphorylated substrates are playing important roles in the cross talk of autophagy and apoptotic cell death at the mitochondria and vesicle nucleation stage of autophagy induction [23,52]. Beclin1 (ATG6) and ULK1 (ATG1) are both involved in autophagosome formation and are direct substrates of AKT [63,68]. AKT is also known to phosphorylate B cell lymphoma (Bcl)-Associated Death promoter (BAD) and triggers the mitochondrial activation of Bcl-2, thereby preventing the release of cytochrome c from mitochondria. Bcl-2 is an anti-apoptotic protein that acts as a major effector of AKT signaling and maintains mitochondrial outer membrane potential to modulate the cell survival, in part by inhibiting Beclin1-dependent autophagy [56]. Consistently, autophagy induced by suberoyanilidehydroxamic acid inhibited AKT and up regulated Beclin1 [69]. The combined inhibition of PI3K and mTOR, activates autophagy without activating AKT (primarily in PTEN [phosphatase and tensin homolog] mutant cells) [70]. The autophagy protein ATG12 associates with anti-apoptotic Bcl-2 family members to promote mitochondrial-mediated apoptosis [64]. Anti-apoptotic protein Bcl-2 is shown to inhibit Beclin 1-dependent autophagy [56].
Involvement of AKT in the regulation of autophagy was further supported by the fact lysosomal accumulation of the AKT-Phafin2 complex is dependent on phosphotidylinositol 3-phosphate and leads to the induction of autophagy [71]. By yeast two-hybrid screening, Phafin2 (EAPF or PLEKHF2), a lysosomal protein with a unique structure of N-terminal PH domain and C-terminal FYVE (Fab 1, YOTB, Vac 1, and EEA1) domain was found to interact with AKT [71]. These conserved motifs place Phafin2 in a family of proteins known to induce caspase-independent apoptosis via the lysosomalmitochondrial pathway [72]. AKT translocates with Phafin2 to the lysosome in a PtdIns (3)P-dependent manner after induction of autophagy. Lysosomal accumulation of the AKT-Phafin2 complex and subsequent induction of autophagy are lysosomal PtdIns (3) P-dependent events, and the formation of this complex at lysosome is a critical step in induction of autophagy via interaction with PtdIns (3)P [71]. These observations also suggest that the regulation of lysosomal localization of AKT, a core anti-apoptotic effector, affects autophagy induction. Therefore, mTORC1 (via TRAF6- p62-mediated ubiquitination) [73] and AKT (via Phafin2-induced autophagy) potentially act as molecular links between autophagy and apoptosis [23].
Additionally, inhibition of mTORC1 (mammalian target of rapamycin complex 1), an effector in the AKT pathway, is known to induce autophagy, the regulation of cell growth, and tumor transformation. mTORC1, a key regulator for autophagy induction consisting of mTOR, Raptor, and mLST8 (mTOR associated protein, LST8 homolog), is shown to be ubiquitinated by the p62-TRAF6 (TNF receptor-associated factor 6) complex at the lysosome The Raptor-mTORC1 is ubiquitinated by the p62- TRAF6 complex to inhibit autophagy presumably at the lysosome [73,74]. mTORC1 is also shown to directly phosphorylate ULK1 (ATG1) and ATG13 [75]. Sch9 kinase (serine/threonine protein kinase 9), the yeast orthologue of mammalian AKT and possibly of ribosomal S6 kinase 1, has been implicated in the regulation of autophagy [76,77]. AKT and TSC (tuberous sclerosis complex) are also localized at lysosome [61,71], suggesting the possibility that they are involved in the regulation of autophagy.
Further support the involvement of AKT as a key molecule for both anti-apoptosis and autophagy regulation, FoxO3, member of the fork head family of transcription factors, is an AKT substrate that regulates the cell death machinery [78]. FoxO3 coordinately activates protein degradation by the autophagic/lysosomal and proteasomal pathways in atrophying muscle cells [79]. Under starvation conditions, FoxO3 activity is shown to be required for a gene expression that induces autophagy in order to mitigate the energy crisis and promote cell survival [80].
Given that mitochondria play an important role in the regulation of apoptosis [6,48-50], these findings support the notion that mitochondria is likely to be an intersection between autophagic cell death and apoptosis [8,81,82]. The observation is also supported by the demonstration by Yoshimori and co-workers that the ER-resident SNARE protein syntaxin 17 (STX17) binds ATG14 and recruits it to the ER-mitochondria contact site, so that the ER-mitochondria contact site is important in autophagosome formation [83].
Conclusion
Autophagy sequesters cytosolic proteins or defective organelles for gross disposal systems of in double or multimembraneautophagic vesicles for degradation and recycling during stress situations such as nutrient starvation in mammalian cells [1,23]. Attention has turned to cross-talk regulation between anti-apoptotic pathways and the induction of autophagy in mammalian cells [6,11,16,17]. Proapoptotic signals such as TRAIL [40], TNF [41], and FADD [42] are also known to induce autophagy. Conversely, anti-apoptotic signaling pathways such as the class I PI3K/AKT/mTOR signaling pathway, suppress autophagy [8]. It was suggested that autophagy may be cytoprotective, at least under conditions of nutrient depletion, and point to an important cross talk between apoptosis and autophagic cell death pathways [24]. Although necrosis is known to be taken place as a consequence of apoptosis [13], it is unclear whether any mechanistic interactions are occurring between the autophagic and necrotic pathways in mammalian cells. It remains to be determined molecular regulatory mechanisms underneath lysosomal degradation of autophagic pathway. In this regard it would be appropriate that autophagic cell death can be defined by which cells are dying with autophagy or cell death associated with autophagy. Thus, in certain scenarios, cell death is associated with autophagy in mammalian cells. Evasion of apoptosis underlies as a pathogenesis of cancer or neoplastic diseases [18,36,84], in which mitochondria is shown to play an important role through cytochrome c release via the disruption of mitochondrial outer membrane potential [6,13].
To further support the potential role of mitochondria as a molecular platform of cross talk between autophagy and apoptosis, numbers of autophagy regulated gene are shown to be controlled by anti-apoptotic molecules such as Bcl-2-BAD-Beclin1 through AKT mediated phosphorylation at the mitochondria [58,63,85,86,]. Indeed, the observation is also supported by the demonstration that autophagosomes form at the ER-mitochondria contact site in mammalian cells [83].
Given the roles of autophagy and apoptosis as underlying mechanisms for cancer [18,35,63,87], the observations support the notion that mitochondria would be the potential molecular platform for counter regulation of autophagic cell death vs. apoptotic cell death.
Further studies are required to precisely clarify the molecular regulation of crosstalk between cell death and survival which maintain the cellular homeostasis in vivo. Autophagic cell death is taking place during the anticancer treatment; therefore, clarification of cross talk between apoptosis and autophagic cell death may provide a platform for developing new cancer treatment modality.
References
- Mizushima N, Komatsu M. Autophagy: renovation of cells and tissues. Cell. 2011; 147: 728-741.
- Virgin HW, Levine B. Autophagy genes in immunity. Nat Immunol. 2009; 10: 461-470.
- Klionsky DJ. The molecular machinery of autophagy: unanswered questions. J Cell Sci. 2005; 118: 7-18.
- Levine B, Mizushima N, Virgin HW. Autophagy in immunity and inflammation. Nature. 2011; 469: 323-335.
- Mizushima N, Levine B, Cuervo AM, Klionsky DJ. Autophagy fights disease through cellular self-digestion. Nature. 2008; 451: 1069-1075.
- Green DR, Galluzzi L, Kroemer G. Mitochondria and the autophagy-inflammation-cell death axis in organismal aging. Science. 2011; 333: 1109-1112.
- Kroemer G, Galluzzi L, Vandenabeele P, Abrams J, Alnemri ES, Baehrecke EH, et al. Classification of cell death: recommendations of the Nomenclature Committee on Cell Death 2009. Cell Death Differ. 2009; 16: 3-11.
- Gozuacik D, Kimchi A. Autophagy as a cell death and tumor suppressor mechanism. Oncogene. 2004; 23: 2891-2906.
- Green DR, Levine B. To be or not to be? How selective autophagy and cell death govern cell fate. Cell. 2014; 157: 65-75.
- Gump JM, Thorburn A. Autophagy and apoptosis: what is the connection? Trends Cell Biol. 2011; 21: 387-392.
- Lum JJ, Bauer DE, Kong M, Harris MH, Li C, Lindsten T, et al. Growth factor regulation of autophagy and cell survival in the absence of apoptosis. Cell. 2005; 120: 237-248.
- Shen HM, Codogno P. Autophagic cell death: Loch Ness monster or endangered species? Autophagy. 2011; 7: 457-465.
- Green D. Means to an End: Apoptosis and Other Cell Death Mechanisms. Cold Spring Harbor Laboratory Press. 2010.
- Tsujimoto Y, Shimizu S. Another way to die: autophagic programmed cell death. Cell Death Differ. 2005; 12 Suppl 2: 1528-1534.
- Thompson CB. Apoptosis in the pathogenesis and treatment of disease. Science. 1995; 267: 1456-1462.
- Kroemer G, Levine B. Autophagic cell death: the story of a misnomer. Nat Rev Mol Cell Biol. 2008; 9: 1004-1010.
- Levine B, Yuan J. Autophagy in cell death: an innocent convict? J Clin Invest. 2005; 115: 2679-2688.
- Mathew R, Karantza-Wadsworth V, White E. Role of autophagy in cancer. Nat Rev Cancer. 2007; 7: 961-967.
- Ryter SW, Mizumura K, Choi AM. The impact of autophagy on cell death modalities. Int J Cell Biol. 2014; 2014: 502676.
- Golstein P, Kroemer G. Cell death by necrosis: towards a molecular definition. Trends Biochem Sci. 2007; 32: 37-43.
- Lu JV, Walsh CM. Programmed necrosis and autophagy in immune function. Immunol Rev. 2012; 249: 205-217.
- Pasparakis M, Vandenabeele P. Necroptosis and its role in inflammation. Nature. 2015; 517: 311-320.
- Noguchi M, Hirata N, Suizu F. The links between AKT and two intracellular proteolytic cascades: ubiquitination and autophagy. Biochim Biophys Acta. 2014; 1846: 342-352.
- Boya P, González-Polo RA, Casares N, Perfettini JL, Dessen P, Larochette N, et al. Inhibition of macroautophagy triggers apoptosis. Mol Cell Biol. 2005; 25: 1025-1040.
- Giusti C, Luciani MF, Golstein P. A second signal for autophagic cell death? Autophagy. 2010; 6: 823-824.
- Su M, Mei Y, Sinha S. Role of the Crosstalk between Autophagy and Apoptosis in Cancer. J Oncol. 2013; 2013: 102735.
- Debnath J, Baehrecke EH, Kroemer G. Does autophagy contribute to cell death? Autophagy. 2005; 1: 66-74.
- Kroemer G, Jäättelä M. Lysosomes and autophagy in cell death control. Nat Rev Cancer. 2005; 5: 886-897.
- Klionsky DJ, Abdalla FC, Abeliovich H, Abraham RT, Acevedo-Arozena A, Adeli K, et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy. 2012; 8: 445-544.
- Klionsky DJ, Abeliovich H, Agostinis P, Agrawal DK, Aliev G, Askew DS, et al. Guidelines for the use and interpretation of assays for monitoring autophagy in higher eukaryotes. Autophagy. 2008; 4: 151-175.
- Mizushima N. Methods for monitoring autophagy. Int J Biochem Cell Biol. 2004; 36: 2491-2502.
- Mizushima N, Yoshimori T, Levine B. Methods in mammalian autophagy research. Cell. 2010; 140: 313-326.
- Mizushima N, Yoshimori T. How to interpret LC3 immunoblotting. Autophagy. 2007; 3: 542-545.
- Komatsu M, Waguri S, Koike M, Sou YS, Ueno T, Hara T, et al. Homeostatic levels of p62 control cytoplasmic inclusion body formation in autophagy-deficient mice. Cell. 2007; 131: 1149-1163.
- Mathew R, Karp CM, Beaudoin B, Vuong N, Chen G, Chen HY, et al. Autophagy suppresses tumorigenesis through elimination of p62. Cell. 2009; 137: 1062-1075.
- Altman BJ, Rathmell JC. Metabolic stress in autophagy and cell death pathways. Cold Spring Harb Perspect Biol. 2012; 4: a008763.
- Ohsumi Y. Molecular dissection of autophagy: two ubiquitin-like systems. Nat Rev Mol Cell Biol. 2001; 2: 211-216.
- Carmona-Gutierrez D, Eisenberg T, Büttner S, Meisinger C, Kroemer G, Madeo F. Apoptosis in yeast: triggers, pathways, subroutines. Cell Death Differ. 2010; 17: 763-773.
- Edinger AL, Thompson CB. Death by design: apoptosis, necrosis and autophagy. Curr Opin Cell Biol. 2004; 16: 663-669.
- Mills KR, Reginato M, Debnath J, Queenan B, Brugge JS. Tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) is required for induction of autophagy during lumen formation in vitro. Proc Natl AcadSci U S A. 2004; 101: 3438-3443.
- Prins J, Ledgerwood E, Ameloot P, Vandenabeele P, Faraco P, Bright N, et al. ().Tumour necrosis factor induced autophagy and mitochondrial morphological abnormalities are mediated by TNFR-I and/or TNFR-II and do not invariably lead to cell death. Biochem Soc Trans. 1998; 26: S314.
- Thorburn J, Moore F, Rao A, Barclay WW, Thomas LR, Grant KW, et al. Selective inactivation of a Fas-associated death domain protein (FADD)-dependent apoptosis and autophagy pathway in immortal epithelial cells. MolBiol Cell. 2005; 16: 1189-1199.
- Thornberry NA, Lazebnik Y. Caspases: enemies within. Science. 1998; 281: 1312-1316.
- Yin X, Cao L, Peng Y, Tan Y, Xie M, Kang R, et al. A critical role for UVRAG in apoptosis. Autophagy. 2011; 7: 1242-1244.
- Cho DH, Jo YK, Hwang JJ, Lee YM, Roh SA, Kim JC. Caspase-mediated cleavage of ATG6/Beclin-1 links apoptosis to autophagy in HeLa cells. Cancer Lett. 2009; 274: 95-100.
- Wang Y, Singh R, Massey AC, Kane SS, Kaushik S, Grant T, et al. Loss of macroautophagy promotes or prevents fibroblast apoptosis depending on the death stimulus. J Biol Chem. 2008; 283: 4766-4777.
- Wang C, Youle RJ. The role of mitochondria in apoptosis*. Annu Rev Genet. 2009; 43: 95-118.
- Kennedy SG, Kandel ES, Cross TK, Hay N. Akt/Protein kinase B inhibits cell death by preventing the release of cytochrome c from mitochondria. Mol Cell Biol. 1999; 19: 5800-5810.
- Laine J, Künstle G, Obata T, Sha M, Noguchi M. The protooncogene TCL1 is an Akt kinase coactivator. Mol Cell. 2000; 6: 395-407.
- Vander Heiden MG, Chandel NS, Li XX, Schumacker PT, Colombini M, Thompson CB. Outer mitochondrial membrane permeability can regulate coupled respiration and cell survival. Proc Natl Acad Sci U S A. 2000; 97: 4666-4671.
- Yousefi S, Perozzo R, Schmid I, Ziemiecki A, Schaffner T, Scapozza L, et al. Calpain-mediated cleavage of Atg5 switches autophagy to apoptosis. Nat Cell Biol. 2006; 8: 1124-1132.
- Marquez RT, Xu L. Bcl-2: Beclin 1 complex: multiple, mechanisms regulating autophagy/apoptosis toggle switch. Am J Cancer Res. 2012; 2: 214-221.
- Radoshevich L, Murrow L, Chen N, Fernandez E, Roy S, Fung C, et al. ATG12 conjugation to ATG3 regulates mitochondrial homeostasis and cell death. Cell. 2010; 142: 590-600.
- Gozuacik D, Bialik S, Raveh T, Mitou G, Shohat G, Sabanay H, et al. DAP-kinase is a mediator of endoplasmic reticulum stress-induced caspase activation and autophagic cell death. Cell Death Differ. 2008; 15: 1875-1886.
- Luo S, Garcia-Arencibia M, Zhao R, Puri C, Toh PP, Sadiq O, et al. Bim inhibits autophagy by recruiting Beclin 1 to microtubules. Mol Cell. 2012; 47: 359-370.
- Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N, et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell. 2005; 122: 927-939.
- Sinha S, Levine B. The autophagy effector Beclin 1: a novel BH3-only protein. Oncogene. 2008; 27 Suppl 1: S137-148.
- He C, Klionsky DJ. Regulation mechanisms and signaling pathways of autophagy. Annu Rev Genet. 2009; 43: 67-93.
- Datta SR, Dudek H, Tao X, Masters S, Fu H, Gotoh Y, et al. Akt phosphorylation of BAD couples survival signals to the cell-intrinsic death machinery. Cell. 1997; 91: 231-241.
- Yang E, Zha J, Jockel J, Boise LH, Thompson CB, Korsmeyer SJ. Bad, a heterodimeric partner for Bcl-XL and Bcl-2, displaces Bax and promotes cell death. Cell. 1995; 80: 285-291.
- Menon S, Dibble CC, Talbott G, Hoxhaj G, Valvezan AJ, Takahashi H, et al. Spatial control of the TSC complex integrates insulin and nutrient regulation of mTORC1 at the lysosome. Cell. 2014; 156: 771-785.
- Djavaheri-Mergny M, Maiuri MC, Kroemer G. Cross talk between apoptosis and autophagy by caspase-mediated cleavage of Beclin 1. Oncogene. 2010; 29: 1717-1719.
- Wang RC, Wei Y, An Z, Zou Z, Xiao G, Bhagat G, White M. Akt-mediated regulation of autophagy and tumorigenesis through Beclin 1 phosphorylation. Science. 2012; 338: 956-959.
- Rubinstein AD, Eisenstein M, Ber Y, Bialik S, Kimchi A. The autophagy protein Atg12 associates with antiapoptotic Bcl-2 family members to promote mitochondrial apoptosis. Mol Cell. 2011; 44: 698-709.
- Jin Z, Li Y, Pitti R, Lawrence D, Pham VC, Lill JR, et al. Cullin3-based polyubiquitination and p62-dependent aggregation of caspase-8 mediate extrinsic apoptosis signaling. Cell. 2009; 137: 721-735.
- Boya P, Andreau K, Poncet D, Zamzami N, Perfettini JL, Metivier D, et al. Lysosomal membrane permeabilization induces cell death in a mitochondrion-dependent fashion. J Exp Med. 2003; 197: 1323-1334.
- Du Toit A. Protein degradation: An alternative route for mitochondrial quality control. Nat Rev Mol Cell Biol. 2014; 15: 150-151.
- Bach M, Larance M, James DE, Ramm G. The serine/threonine kinase ULK1 is a target of multiple phosphorylation events. Biochem J. 2011; 440: 283-291.
- Cao Q, Yu C, Xue R, Hsueh W, Pan P, Chen Z, et al. Autophagy induced by suberoylanilide hydroxamic acid in Hela S3 cells involves inhibition of protein kinase B and up-regulation of Beclin 1. Int J Biochem Cell Biol. 2008; 40: 272-283.
- Fan QW, Cheng C, Hackett C, Feldman M, Houseman BT, Nicolaides T, et al. Akt and autophagy cooperate to promote survival of drug-resistant glioma. Sci Signal. 2010; 3: ra81.
- Matsuda-Lennikov M, Suizu F, Hirata N, Hashimoto M, Kimura K, Nagamine T, et al. Lysosomal interaction of Akt with Phafin2: a critical step in the induction of autophagy. PLoS One. 2014; 9: e79795.
- Chen W, Li N, Chen T, Han Y, Li C, Wang Y, et al. The lysosome-associated apoptosis-inducing protein containing the pleckstrin homology (PH) and FYVE domains (LAPF), representative of a novel family of PH and FYVE domain-containing proteins, induces caspase-independent apoptosis via the lysosomal-mitochondrial pathway. J Biol Chem. 2005; 280: 40985-40995.
- Linares JF, Duran A, Yajima T, Pasparakis M, Moscat J, Diaz-Meco MT. K63 polyubiquitination and activation of mTOR by the p62-TRAF6 complex in nutrient-activated cells. Mol Cell. 2013; 51: 283-296.
- Poüs C, Codogno P. Lysosome positioning coordinates mTORC1 activity and autophagy. Nat Cell Biol. 2011; 13: 342-344.
- Wong PM, Puente C, Ganley IG, Jiang X. The ULK1 complex: sensing nutrient signals for autophagy activation. Autophagy. 2013; 9: 124-137.
- Kaeberlein M, Powers RW 3rd, Steffen KK, Westman EA, Hu D, Dang N, et al. Regulation of yeast replicative life span by TOR and Sch9 in response to nutrients. Science. 2005; 310: 1193-1196.
- Madia F, Wei M, Yuan V, Hu J, Gattazzo C, Pham P, et al. Oncogene homologue Sch9 promotes age-dependent mutations by a superoxide and Rev1/Polzeta-dependent mechanism. J Cell Biol. 2009; 186: 509-523.
- Brunet A, Bonni A, Zigmond MJ, Lin MZ, Juo P, Hu LS, et al. Akt promotes cell survival by phosphorylating and inhibiting a Forkhead transcription factor. Cell. 1999; 96: 857-868.
- Zhao J, Brault JJ, Schild A, Cao P, Sandri M, Schiaffino S, et al. FoxO3 coordinately activates protein degradation by the autophagic/lysosomal and proteasomal pathways in atrophying muscle cells. Cell metabolism 2007; 6: 472-483.
- Warr MR, Binnewies M, Flach J, Reynaud D, Garg T, Malhotra R, et al. FOXO3A directs a protective autophagy program in haematopoietic stem cells. Nature. 2013; 494: 323-327.
- Okamoto K, Kondo-Okamoto N. Mitochondria and autophagy: critical interplay between the two homeostats. Biochim Biophys Acta. 2012; 1820: 595-600.
- Randow F, Youle RJ. Self and nonself: how autophagy targets mitochondria and bacteria. Cell Host Microbe. 2014; 15: 403-411.
- Hamasaki M, Furuta N, Matsuda A, Nezu A, Yamamoto A, Fujita N, et al. Autophagosomes form at ER-mitochondria contact sites. Nature. 2013; 495: 389-393.
- Noguchi M, Ropars V, Roumestand C, Suizu F. Proto-oncogene TCL1: more than just a coactivator for Akt. FASEB J. 2007; 21: 2273-2284.
- Chipuk JE, Moldoveanu T, Llambi F, Parsons MJ, Green DR. The BCL-2 family reunion. Mol Cell. 2010; 37: 299-310.
- Levine B, Sinha S, Kroemer G. Bcl-2 family members: dual regulators of apoptosis and autophagy. Autophagy. 2008; 4: 600-606.
- Radogna F, Dicato M, Diederich M. Cancer-type-specific crosstalk between autophagy, necroptosis and apoptosis as a pharmacological target. Biochem Pharmacol. 2015; 94: 1-11.